TVB-O supports surface-based partial differential equations using finite element methods via scikit-fem . The PDE backend generates a self-contained implicit Euler FEM solver from a YAML experiment description, supporting triangle and tetrahedral meshes, Dirichlet boundary conditions, and constant or time-dependent source terms.
\[
\partial_t u(t,\mathbf{x}) = D\,\Delta u(t,\mathbf{x}) \quad \text{in } \Omega, \qquad u(t,\mathbf{x}) = 0 \quad \text{on } \partial\Omega
\]
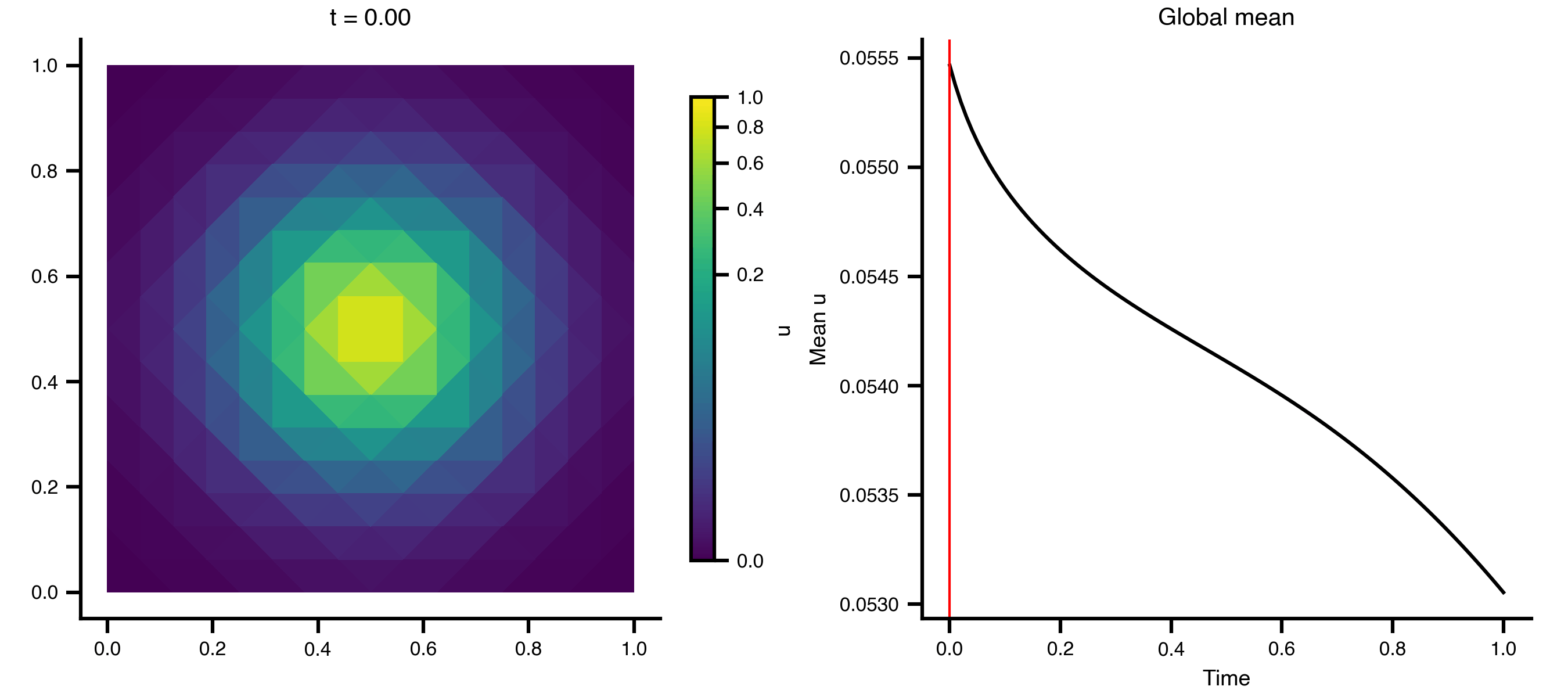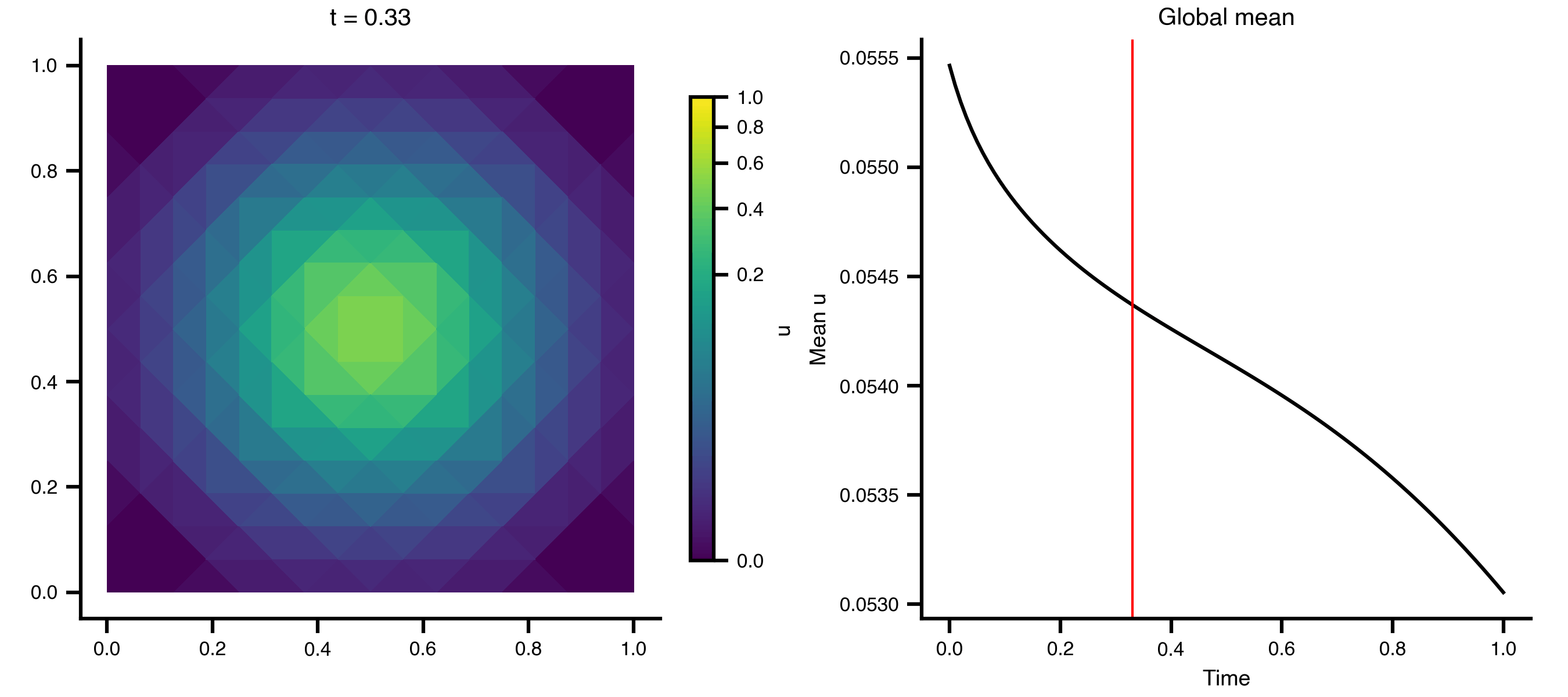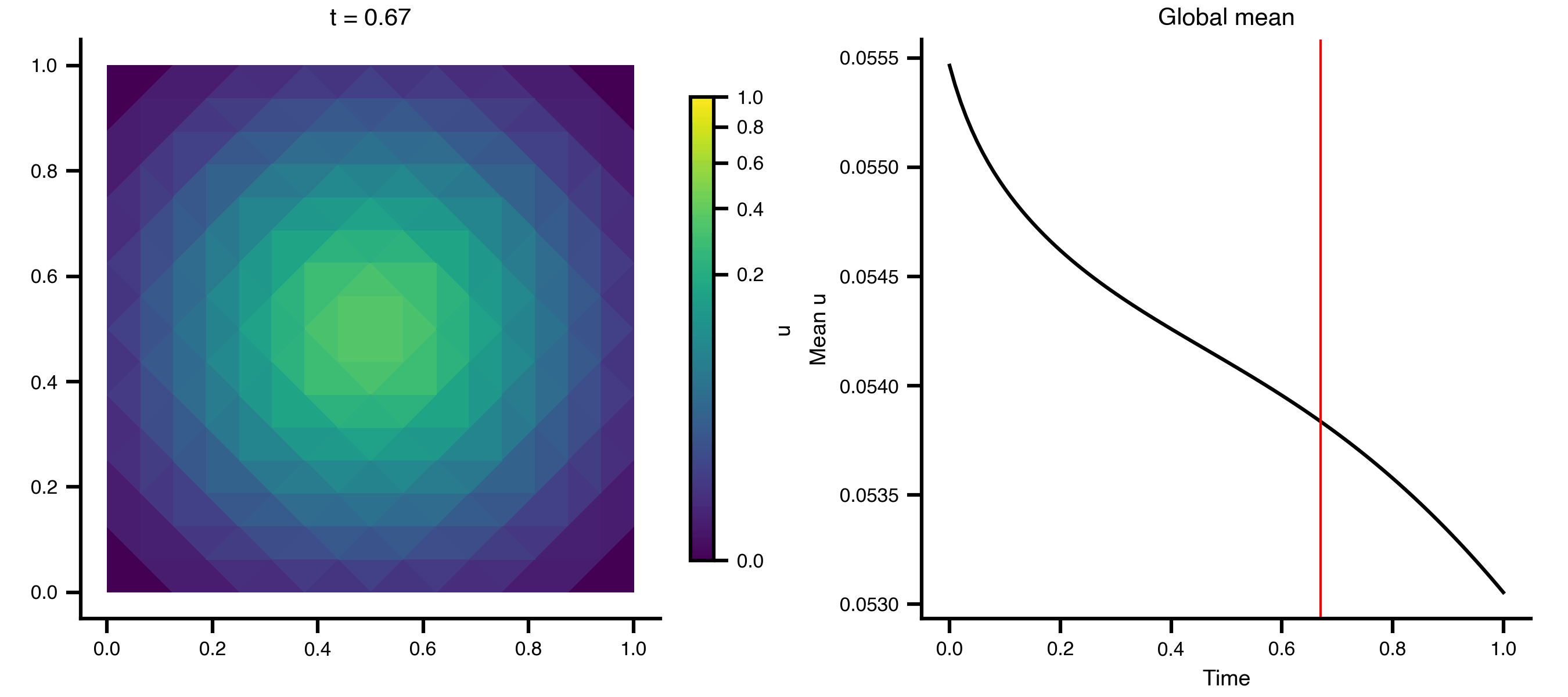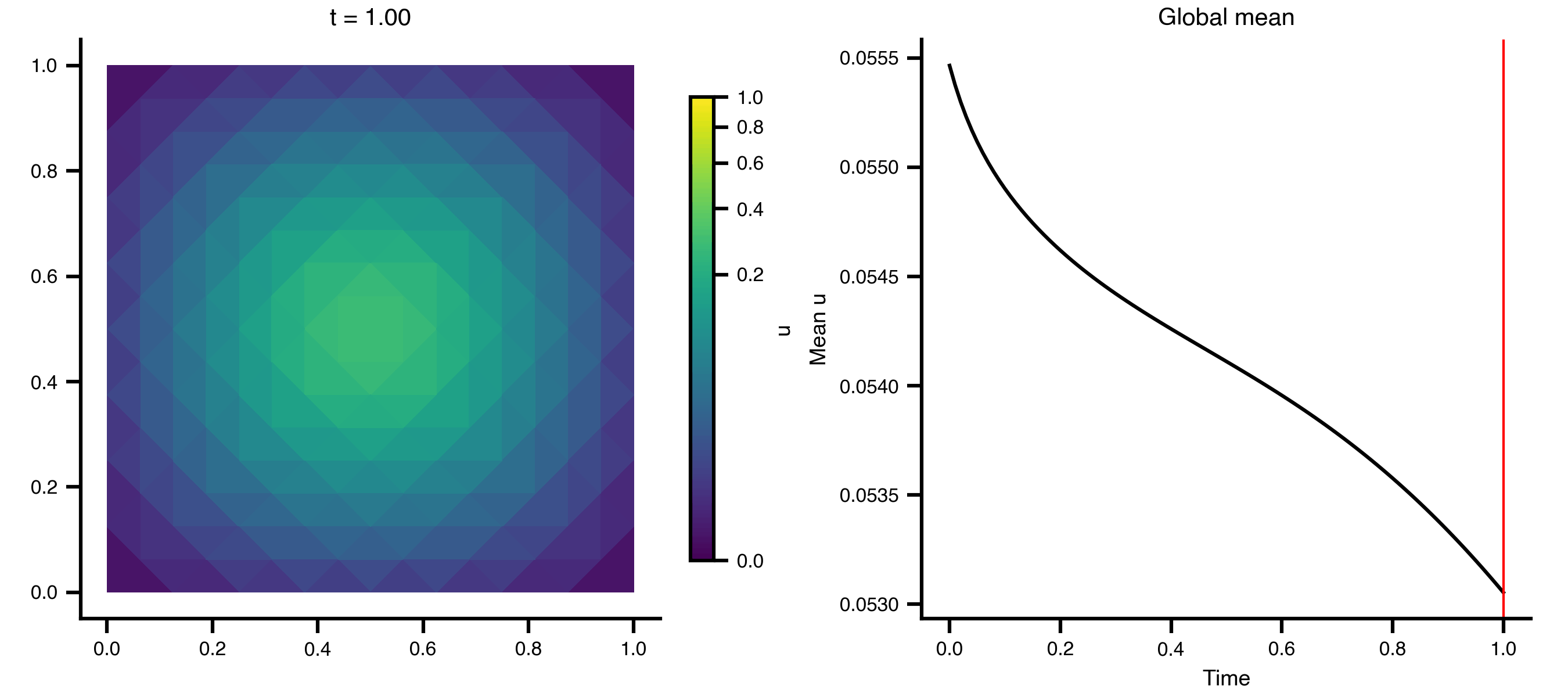
Example 1: Heat Equation on a 2D Grid
A minimal diffusion example on a unit square, using scikit-fem’s built-in mesh generators.
Step 1 — Create a triangle mesh and experiment YAML
import tempfile, os, yamlimport meshioimport numpy as npfrom skfem import MeshTrifrom bsplot import style"tvbo" )# Generate a symmetric unit-square triangulation = MeshTri.init_symmetric().refined(3 )= tempfile.mkdtemp()= os.path.join(tmpdir, "unit_square.msh" )= mesh.p.T, cells= [("triangle" , mesh.t.T)]).write(mesh_path)print (f"Mesh: { mesh. p. shape[1 ]} vertices, { mesh. t. shape[1 ]} triangles" )# Write experiment YAML = {"label" : "Heat equation on unit square" ,"field_dynamics" : {"label" : "Heat / diffusion equation" ,"mesh" : {"label" : "unit_square" ,"element_type" : "triangle" ,"mesh_file" : mesh_path,"parameters" : {"D" : {"name" : "D" , "value" : 0.01 }},"state_variables" : ["name" : "u" ,"label" : "Temperature" ,"initial_value" : 0.0 ,"boundary_conditions" : ["label" : "Zero Dirichlet" ,"bc_type" : "Dirichlet" ,"value" : {"rhs" : "0" },"equation" : {"lhs" : "u_t" , "rhs" : "D * laplacian(u)" },"operators" : ["label" : "Diffusion" , "operator_type" : "laplacian" , "coefficient" : "D" }"solver" : {"label" : "FEM implicit Euler" ,"discretization" : "FEM" ,"time_integrator" : "implicit Euler" ,"dt" : 0.01 ,"integration" : {"duration" : 1.0 },= os.path.join(tmpdir, "pde_heat_2d.yaml" )with open (yaml_path, "w" ) as f:
Mesh: 145 vertices, 256 triangles
Step 2 — Run the simulation
from tvbo import SimulationExperiment= SimulationExperiment.from_file(yaml_path)= exp.execute("pde" )= ns["meta" ]["nodes" ]= nodes[0 ], nodes[1 ]# Gaussian bump initial condition centred at (0.5, 0.5) = np.exp(- ((x - 0.5 )** 2 + (y - 0.5 )** 2 ) / 0.02 )= exp.run("pde" , u0= u0)print (f"TimeSeries: { ts. data. shape} (time, state_var, nodes)" )print (f"Time: { ts. time[0 ]:.2f} to { ts. time[- 1 ]:.2f} , dt= { ts. time[1 ] - ts. time[0 ]:.3f} " )
TimeSeries: (101, 1, 145) (time, state_var, nodes)
Time: 0.00 to 1.00, dt=0.010
Step 3 — Animate the diffusion
import matplotlib.pyplot as pltimport matplotlib.tri as mtrifrom matplotlib.colors import PowerNormfrom matplotlib.animation import PillowWriterfrom IPython.display import Image= mtri.Triangulation(x, y, ns["meta" ]["cells" ].T)= ts.data.shape[0 ]= 0 , float (np.max (ts.data))= PowerNorm(gamma= 0.3 , vmin= vmin, vmax= vmax)= plt.subplots(1 , 2 , figsize= (9 , 4 ), layout= "compressed" )# Add a fixed colorbar (norm is constant across frames) import matplotlib.cm as cm= cm.ScalarMappable(norm= norm, cmap= "viridis" )= ax_mesh, label= "u" , shrink= 0.8 )def update_2d(frame):0 , :]), norm= norm, cmap= "viridis" )f"t = { ts. time[frame]:.2f} " )"equal" )0 , :]).mean(axis= 1 ), color= "black" )= "red" , lw= 1 )"Time" )"Mean u" )"Global mean" )"_output" , exist_ok= True )from matplotlib.animation import FuncAnimation= FuncAnimation(fig, update_2d, frames= n_frames, interval= 100 )"_output/pde_heat_2d.gif" , writer= "pillow" , fps= 10 )
Example 2: Cortical Surface Diffusion
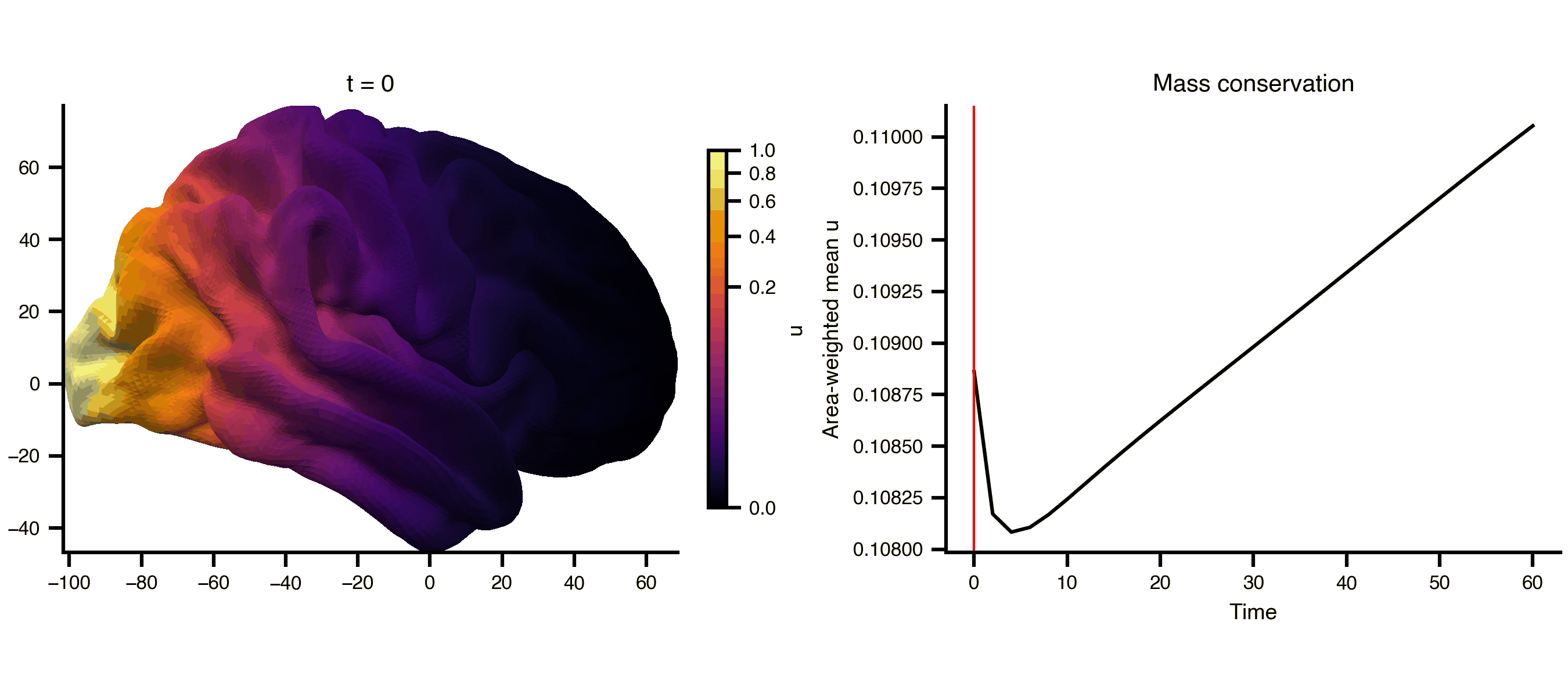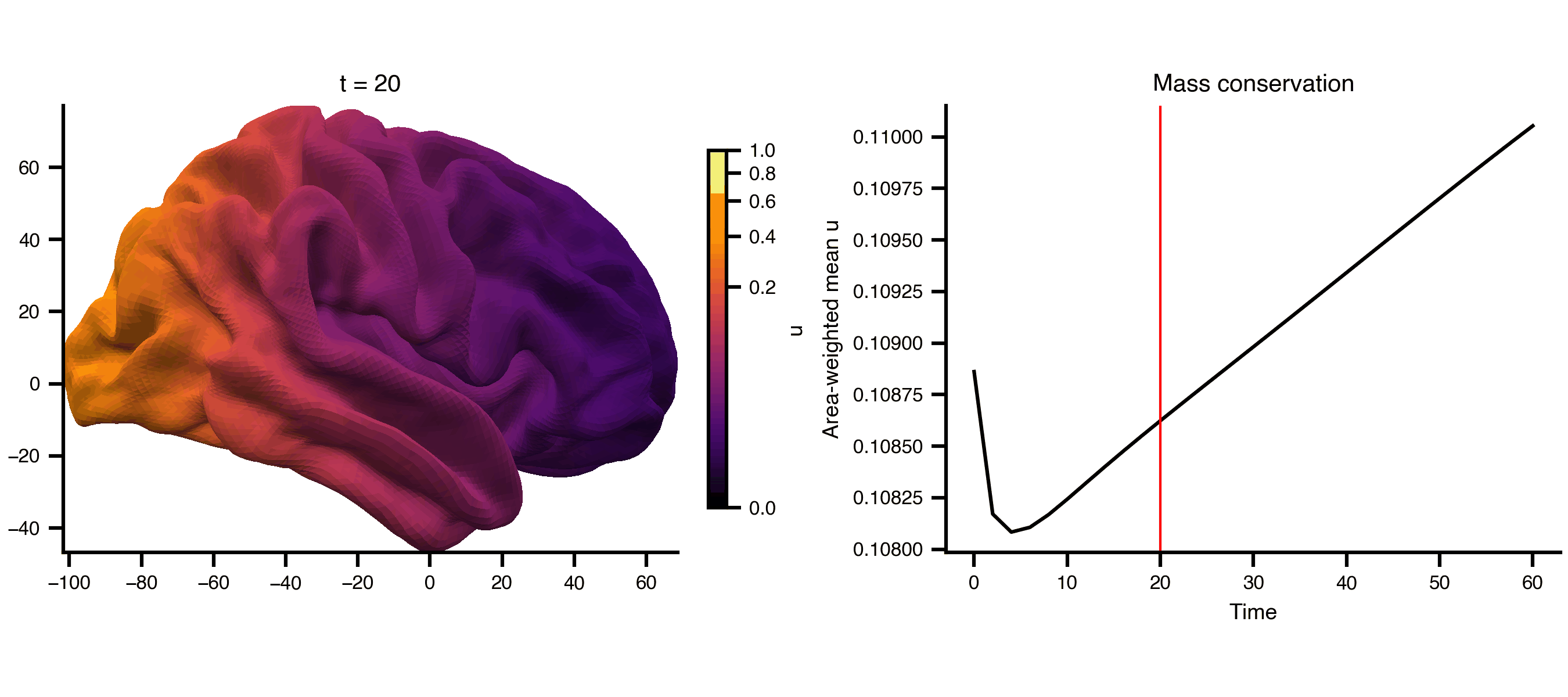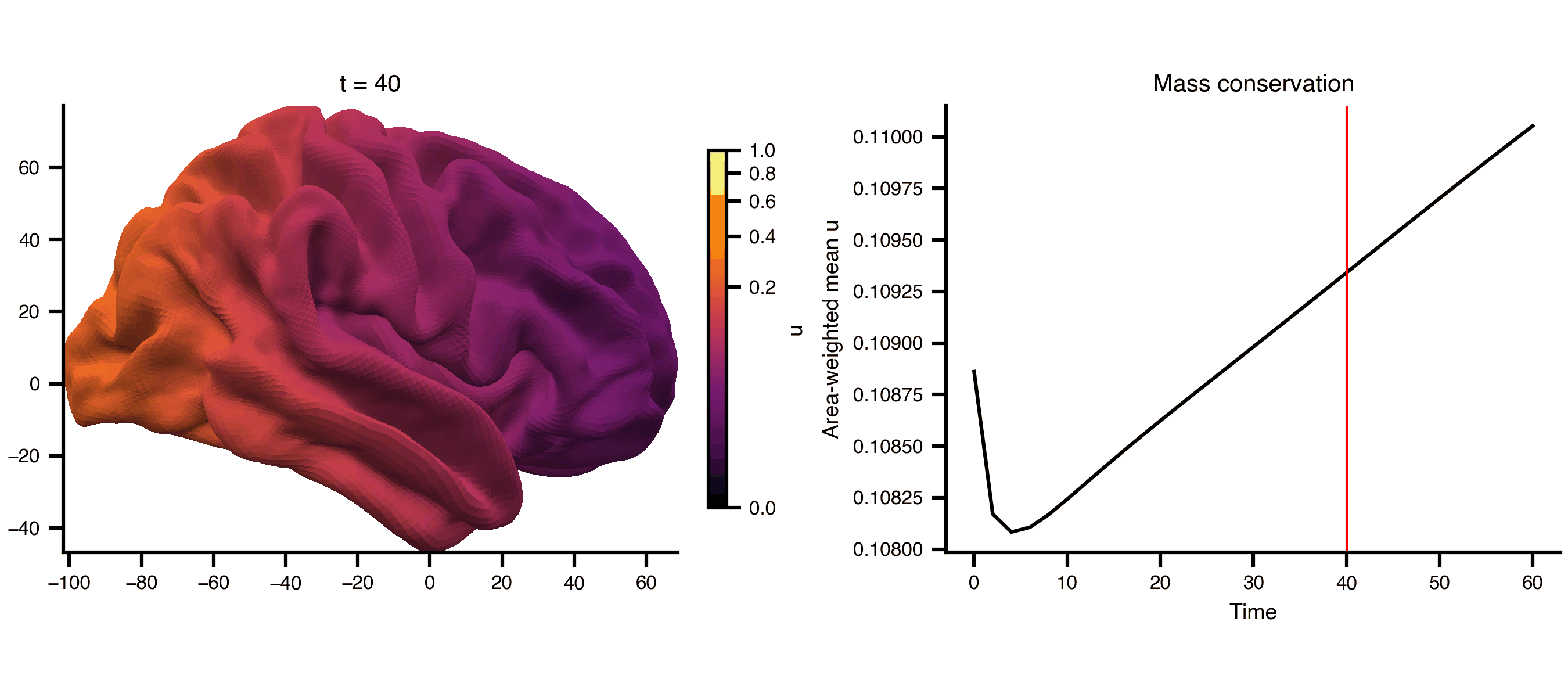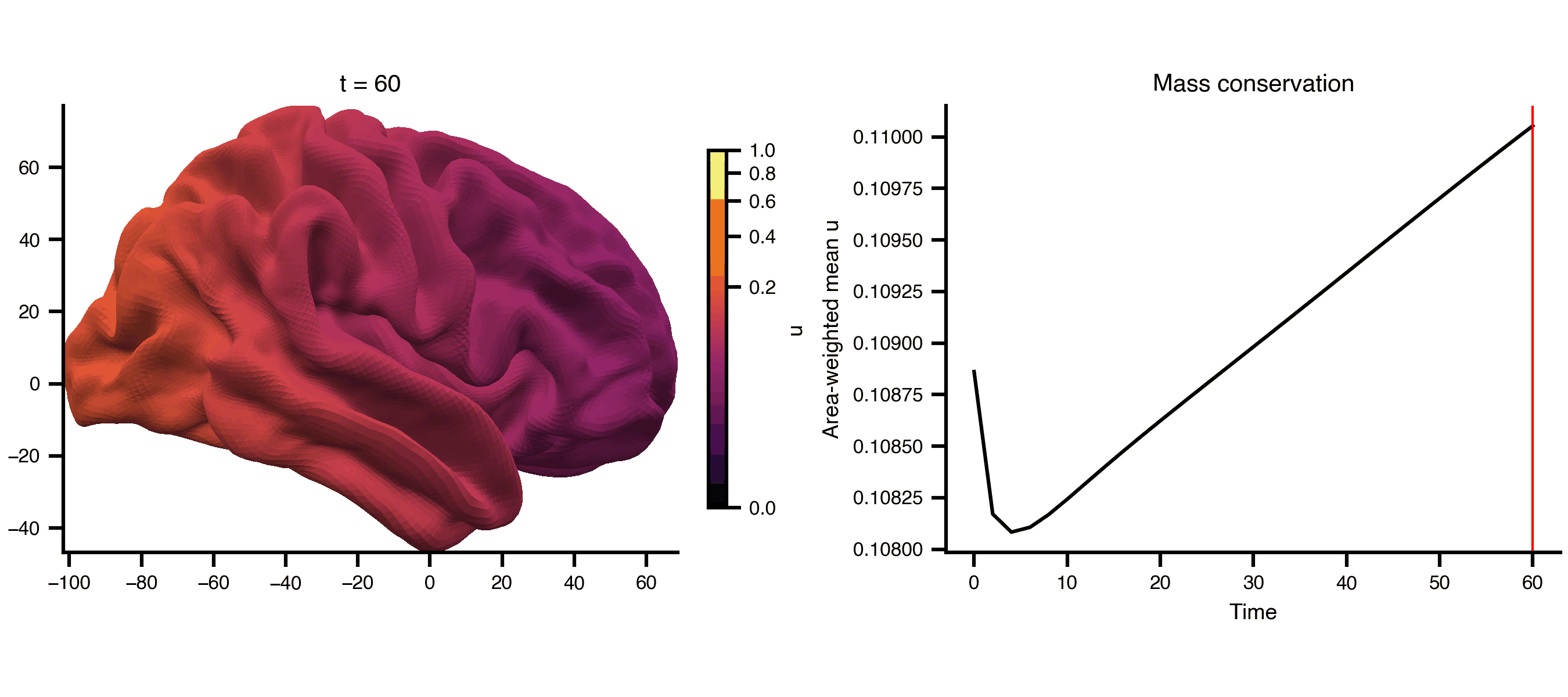
Diffusion on a cortical surface mesh (fsLR 32k), using the mesh_file attribute to reference an external GIFTI file directly. The cortical midthickness surface is a closed manifold (no boundary), so the Dirichlet BC has no effect and total mass is conserved — the field spreads but doesn’t decay.
Setup and run
import templateflow.api as tfaimport nibabel as nib# Get RH midthickness surface from templateflow = str (tfa.get(template= "fsLR" , density= "32k" , suffix= "midthickness" , hemi= "R" , desc= None ))# D=50 gives visible spreading on mm-scale cortical mesh (edge ≈ 1.6 mm) = {'label' : 'Cortical surface diffusion' ,'field_dynamics' : {'label' : 'Heat/diffusion equation' ,'mesh' : {'label' : 'cortex_rh' ,'element_type' : 'triangle' ,'mesh_file' : mesh_gii,'mesh_format' : 'gifti' ,'parameters' : {'D' : {'name' : 'D' , 'value' : 50.0 }},'state_variables' : [{'name' : 'u' , 'label' : 'u' ,'initial_value' : 0.0 ,'boundary_conditions' : [{'label' : 'Zero Dirichlet' , 'bc_type' : 'Dirichlet' , 'value' : {'rhs' : '0' }}],'equation' : {'lhs' : 'u_t' , 'rhs' : 'D * laplacian(u)' },'operators' : [{'label' : 'Diffusion' , 'operator_type' : 'laplacian' , 'coefficient' : 'D' }],'solver' : {'label' : 'FEM IE' , 'discretization' : 'FEM' , 'time_integrator' : 'implicit Euler' , 'dt' : 2.0 },'integration' : {'duration' : 60 },= tempfile.mkdtemp()= os.path.join(tmpdir2, 'pde_cortex.yaml' )with open (yaml_path2, 'w' ) as f:= SimulationExperiment.from_file(yaml_path2)# Load mesh for initial condition = nib.load(mesh_gii)= gi.darrays[0 ].data# Gaussian centred at the most posterior vertex (occipital pole), σ ≈ 45 mm = np.argmin(vertices[:, 1 ])= vertices[seed_idx]= np.sum ((vertices - seed)** 2 , axis= 1 )= np.exp(- dist_sq / 2000 )= exp2.run("pde" , u0= u0)print (f"Cortical PDE: { ts2. data. shape} , { vertices. shape[0 ]} vertices" )
Cortical PDE: (31, 1, 32492), 32492 vertices
Animate with bsplot
from matplotlib.colors import PowerNormfrom bsplot.surface import plot_surffrom bsplot.animate import animate_axes= ts2.data.shape[0 ]= float (np.max (ts2.data))= PowerNorm(gamma= 0.3 , vmin= 0 , vmax= vmax2)= plt.subplots(1 , 2 , figsize= (9 , 4 ), layout= "compressed" )# Add a fixed colorbar import matplotlib.cm as cm= cm.ScalarMappable(norm= norm2, cmap= "inferno" )= ax_surf, label= "u" , shrink= 0.8 )# Area-weighted mean (conserved on closed surface), not arithmetic mean = gi.darrays[1 ].data= vertices[faces[:, 0 ]], vertices[faces[:, 1 ]], vertices[faces[:, 2 ]]= 0.5 * np.linalg.norm(np.cross(v1 - v0, v2 - v0), axis= 1 )= np.zeros(len (vertices))for k in range (3 ):/ 3 )= np.array([* np.asarray(ts2.data[t, 0 , :])).sum () / vertex_areas.sum ()for t in range (n_frames)def cortex_update(frame, ax, axis_idx, context):if axis_idx == 0 := np.asarray(context["ts" ].data[frame, 0 , :])"gii" ], overlay= overlay,= "rh" , view= "lateral" , ax= ax,= "inferno" , norm= context["norm" ],f"t = { context['ts' ]. time[frame]:.0f} " )return []else :"ts" ].time, context["mean" ], color= "black" )"ts" ].time[frame], color= "red" , lw= 1 )"Time" )"Area-weighted mean u" )"Mass conservation" )return []= {"ts" : ts2, "gii" : gi, "norm" : norm2, "mean" : global_mean}= animate_axes(fig2, [ax_surf, ax_ts], n_frames= n_frames,= cortex_update, context= ctx, interval= 200 )"_output/pde_cortex.gif" , writer= "pillow" , fps= 6 )
Numerical details
The backward-Euler FEM update assembles mass (\(M\) ) and stiffness (\(K\) ) matrices over linear triangular elements and solves, at each step:
\[
(M + \Delta t \, D \, K) \, \mathbf{u}^{n+1} = M \, \mathbf{u}^{n} + \Delta t \, M \, \mathbf{f}^{n}
\]
Mesh Support
The Mesh class supports two ways to reference mesh geometry:
mesh_filePath to external mesh file (GIFTI, VTK, MSH, FreeSurfer, etc.)
mesh_formatExplicit format override (gifti, freesurfer, meshio). Auto-detected from extension if omitted
dataLocationLegacy attribute — still works, supports prefix syntax (gifti:path/to/file.gii)
Supported mesh formats:
GIFTI
.gii, .surf.giinibabel
FreeSurfer
.pial, .white, .inflatednibabel
Gmsh
.mshmeshio
VTK
.vtk, .vtumeshio
Wavefront OBJ
.objmeshio
PLY
.plymeshio
Generated Code
The PDE backend generates a self-contained scikit-fem solver from the YAML specification:
= exp.render_code("pde" )print (code[:500 ])The generated module contains:
_load_mesh()Load GIFTI, FreeSurfer, or meshio mesh formats
build()Assemble mass and stiffness matrices, return solver closures
solve_pde()Implicit Euler timestepping with optional source terms
visualize()Matplotlib field plot via scikit-fem
Schema Classes
PDETop-level PDE problem definition
SpatialDomainCoordinate space and geometry
MeshTriangle/tetrahedron element mesh with mesh_file/mesh_format
FieldStateVariableSpatially distributed state variable
DifferentialOperatorGradient, divergence, laplacian, curl
BoundaryConditionDirichlet, Neumann, Robin, Periodic
PDESolverFEM/FDM/FVM discretization config