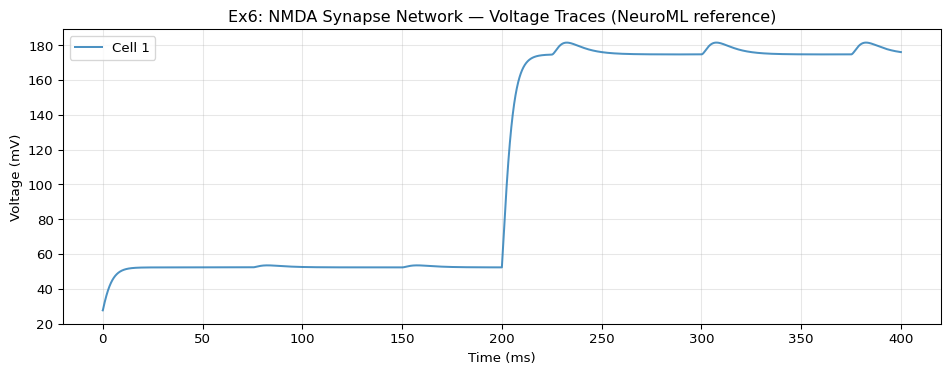
Model: NMDA Synapse
A passive cell (with morphology and biophysics) receives NMDA synapse input from a spike generator, plus a DC current injection. Demonstrates the cell type with channelDensity and blockingPlasticSynapse with Mg²⁺ block.
Reference: NeuroML2 LEMS_NML2_Ex6_NMDA.xml
1. Define Network in TVBO
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NeuroML Ex6: NMDA Synapse" network: dynamics: passiveCell: name: passiveCell iri: neuroml:cell parameters: diameter: {value: 17.841242} specificCapacitance: {value: 1.0, unit: uF_per_cm2} initMembPotential: {value: -65, unit: mV} spikeThresh: {value: -20, unit: mV} resistivity: {value: 0.03, unit: kohm_cm} components: passiveChan: name: passiveChan iri: neuroml:ionChannelHH parameters: conductance: {value: 10, unit: pS} condDensity: {value: 0.0003, unit: S_per_cm2} erev: {value: -54.3, unit: mV} ion: {description: non_specific} spikeGen75ms: name: spikeGen75ms iri: neuroml:spikeGenerator parameters: period: {value: 75, unit: ms} nmdaSyn1: name: nmdaSyn1 iri: neuroml:blockingPlasticSynapse parameters: gbase: {value: 0.5, unit: nS} erev: {value: 0, unit: mV} tauRise: {value: 2, unit: ms} tauDecay: {value: 8, unit: ms} modes: blockMechanism: name: blockMechanism iri: neuroml:voltageConcDepBlockMechanism parameters: species: {description: mg} blockConcentration: {value: 1.2, unit: mmol_per_m3} scalingConc: {value: 1.920544, unit: mmol_per_m3} scalingVolt: {value: 16.129, unit: mV} pulseGen2: name: pulseGen2 iri: neuroml:pulseGenerator parameters: delay: {value: 200, unit: ms} duration: {value: 10000, unit: ms} amplitude: {value: 0.065, unit: nA} nodes: - {id: 0, dynamics: passiveCell} - {id: 10, dynamics: spikeGen75ms} - {id: 100, dynamics: pulseGen2} edges: - {source: 10, target: 0, coupling: nmdaSyn1} - {source: 100, target: 0} integration: method: euler step_size: 0.01 duration: 400.0 time_scale: ms """ )print (f"TVBO model: { exp. dynamics. name if exp. dynamics else "network" } " )
2. Render LEMS XML
= exp.render("lems" )print (xml[:3000 ])
<Lems>
<Target component="sim_NeuroML_Ex6__NMDA_Synapse"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Inputs.xml"/>
<Include file="Simulation.xml"/>
<ionChannelHH id="passiveChan" conductance="10 pS"/>
<cell id="passiveCell">
<morphology id="passiveCell_morphology">
<segment id="0" name="Soma">
<proximal x="0.0" y="0.0" z="0.0" diameter="17.841242"/>
<distal x="0.0" y="0.0" z="0.0" diameter="17.841242"/>
</segment>
<segmentGroup id="all">
<member segment="0"/>
</segmentGroup>
<segmentGroup id="soma_group">
<member segment="0"/>
</segmentGroup>
</morphology>
<biophysicalProperties id="biophys">
<membraneProperties>
<channelDensity condDensity="0.0003 S_per_cm2" id="passiveChan_all" ionChannel="passiveChan" erev="-54.3 mV" ion="non_specific"/>
<specificCapacitance value="1 uF_per_cm2"/>
<initMembPotential value="-65 mV"/>
<spikeThresh value="-20 mV"/>
</membraneProperties>
<intracellularProperties>
<resistivity value="0.03 kohm_cm"/>
</intracellularProperties>
</biophysicalProperties>
</cell>
<spikeGenerator id="spikeGen75ms" period="75.0 ms"/>
<pulseGenerator id="pulseGen2" amplitude="0.065 nA" delay="200.0 ms" duration="10000.0 ms"/>
<blockingPlasticSynapse id="nmdaSyn1" erev="0 mV" gbase="0.5 nS" tauDecay="8 ms" tauRise="2 ms">
<blockMechanism type="voltageConcDepBlockMechanism" blockConcentration="1.2 mM" scalingConc="1.920544 mM" scalingVolt="16.129 mV" species="mg"/>
</blockingPlasticSynapse>
<network id="net1">
<population id="passiveCell_pop" component="passiveCell" size="1"/>
<population id="spikeGen75ms_pop" component="spikeGen75ms" size="1"/>
<synapticConnection from="spikeGen75ms_pop[0]" to="passiveCell_pop[0]" synapse="nmdaSyn1" destination="synapses"/>
<explicitInput target="passiveCell_pop[0]" input="pulseGen2" destination="synapses"/>
</network>
<Simulation id="sim_NeuroML_Ex6__NMDA_Synapse" length="400.0ms" step="0.01ms" target="net1">
<OutputFile id="of0" fileName="results/passiveCell.dat">
<OutputColumn id="passiveCell_pop_0_v" quantity="passiveCell_pop[0]/v"/>
</OutputFile>
</Simulation>
</Lems>
3. Run Reference
from tvbo.adapters.neuroml import run_lems_example= run_lems_example("LEMS_NML2_Ex6_NMDA.xml" )for name, arr in ref_outputs.items():print (f" { name} : shape= { arr. shape} " )
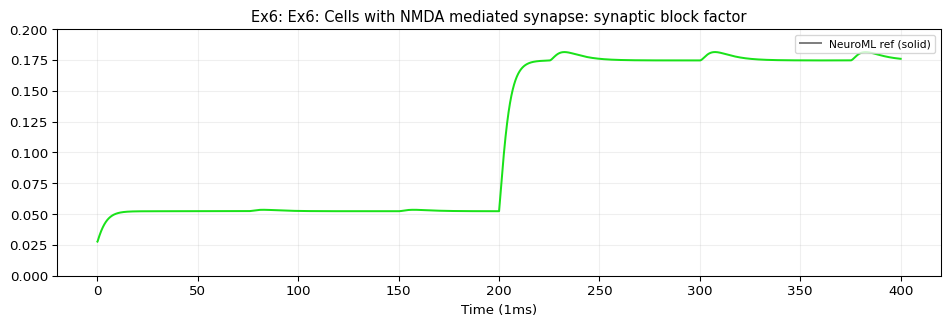
ex6_block.dat: shape=(40001, 2)
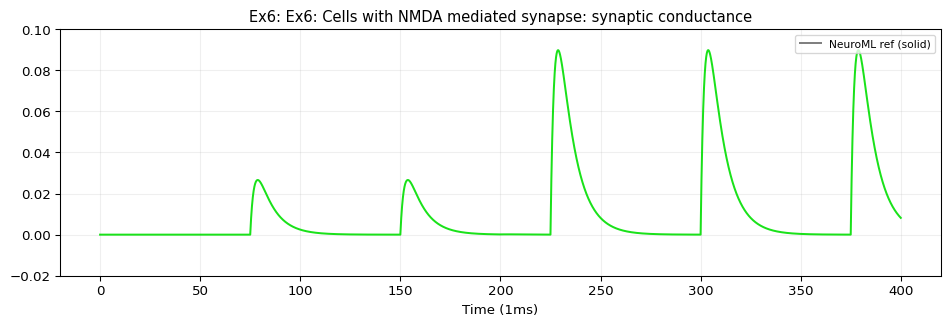
ex6_g.dat: shape=(40001, 2)
ex6_v.dat: shape=(40001, 2)
4. Run TVBO
= exp.run("neuroml" )= result.integration.dataprint (f"TVBO: { da. dims} , shape= { da. shape} " )
TVBO: ('time', 'quantity'), shape=(40001, 1)
5. Compare & Plot
from tvbo.adapters.neuroml import plot_lems_comparison"LEMS_NML2_Ex6_NMDA.xml" , ref_outputs, result.integration.data, title_prefix= "Ex6" )