Model: Tissue with Varying Temperature
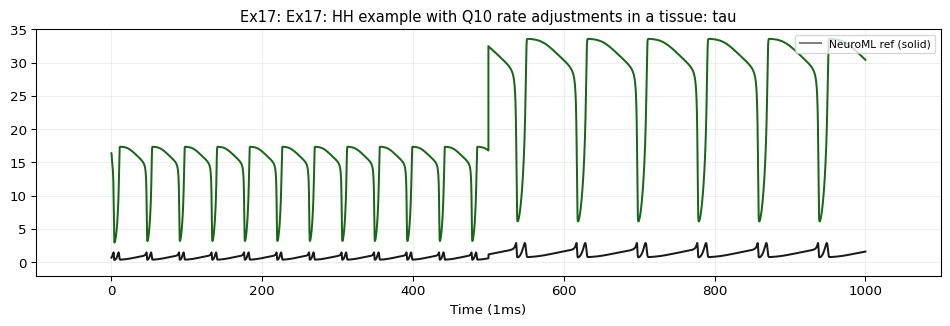
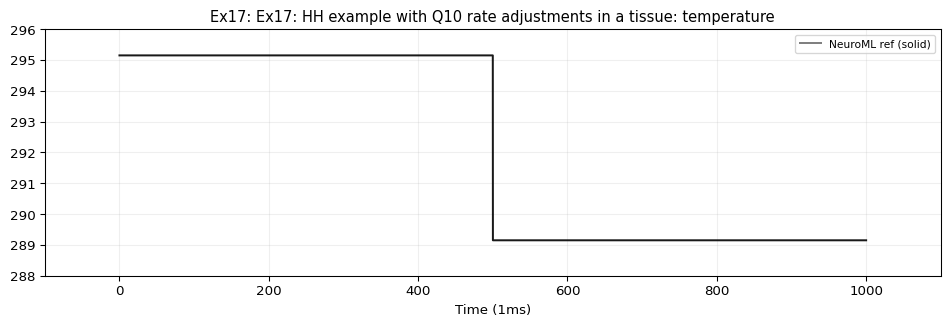
Demonstrates tissueWithVaryingTemperature — an HH cell with Q10-scaled gating rates inside a tissue that changes temperature from 22°C to 16°C at \(t=500\,\text{ms}\) .
\[C\frac{dv}{dt} = -g_{Na}m^3h(v - E_{Na}) - g_K n^4(v - E_K) - g_L(v - E_L) + I_{\text{ext}}\]
Gate rates are scaled by \(Q_{10}(T)\) : \[Q_{10}(T) = 3^{(T - 32)/10}\]
\(T = 22°C\) for \(t < 500\,\text{ms}\) : \(Q_{10} = 3^{-1} = 1/3\) \(T = 16°C\) for \(t \ge 500\,\text{ms}\) : \(Q_{10} = 3^{-1.6} \approx 0.1724\)
1. Define in TVBO
from tvbo import SimulationExperiment# Uses standard NeuroML types: pointCellCondBased + ionChannelHH + gateHHrates # Q10 temperature scaling handled via q10Settings on each gate # Tissue wrapper changes temperature from 22°C to 16°C at 500ms = SimulationExperiment.from_string(""" label: "NeuroML Ex17: HH Tissue with Q10" dynamics: name: HH_Tissue_Q10 iri: neuroml:pointCellCondBased description: > HH cell with Q10 temperature scaling in a tissue that changes temperature from 22°C to 16°C at t=500ms. parameters: C: { value: 10, unit: pF } v0: { value: -65, unit: mV } thresh: { value: 20, unit: mV } pulse_delay: { value: 0, unit: ms } pulse_duration: { value: 1000, unit: ms } I_amp: { value: 0.08, unit: nA } tissue_startTemperature: { value: 22, unit: degC } tissue_endTemperature: { value: 16, unit: degC } tissue_changeTime: { value: 500, unit: ms } components: passive: name: passive iri: neuroml:ionChannelPassive parameters: conductance: { value: 10, unit: pS } number: { value: 300 } erev: { value: -54.3, unit: mV } na: name: na iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 120000 } erev: { value: 50, unit: mV } components: m: name: m iri: neuroml:gateHHrates parameters: instances: { value: 3 } components: q10Settings: name: q10Settings iri: neuroml:q10ExpTemp parameters: q10Factor: { value: 3 } experimentalTemp: { value: 32, unit: degC } forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -40, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 4, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -18, unit: mV } h: name: h iri: neuroml:gateHHrates parameters: instances: { value: 1 } components: q10Settings: name: q10Settings iri: neuroml:q10ExpTemp parameters: q10Factor: { value: 3 } experimentalTemp: { value: 32, unit: degC } forwardRate: name: forwardRate iri: neuroml:HHExpRate parameters: rate: { value: 0.07, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -20, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHSigmoidRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -35, unit: mV } scale: { value: 10, unit: mV } k: name: k iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 36000 } erev: { value: -77, unit: mV } components: n: name: n iri: neuroml:gateHHrates parameters: instances: { value: 4 } components: q10Settings: name: q10Settings iri: neuroml:q10ExpTemp parameters: q10Factor: { value: 3 } experimentalTemp: { value: 32, unit: degC } forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 0.1, unit: per_ms } midpoint: { value: -55, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 0.125, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -80, unit: mV } integration: step_size: 0.01 duration: 1000.0 time_scale: ms """ )print (f"Model: { exp. dynamics. name if exp. dynamics else 'network' } " )print (f"SVs: { list (exp.dynamics.state_variables.keys())} " )
Model: HH_Tissue_Q10
SVs: []
2. Render LEMS XML
= exp.render("lems" )print (xml[:1200 ])
<Lems>
<Target component="sim_NeuroML_Ex17__HH_Tissue_with_Q10"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Inputs.xml"/>
<Include file="Simulation.xml"/>
<ionChannelHH id="k" conductance="10 pS">
<gateHHrates id="n" instances="4">
<forwardRate type="HHExpLinearRate" midpoint="-55 mV" rate="0.1 per_ms" scale="10 mV"/>
<q10Settings type="q10ExpTemp" experimentalTemp="32 degC" q10Factor="3"/>
<reverseRate type="HHExpRate" midpoint="-65 mV" rate="0.125 per_ms" scale="-80 mV"/>
</gateHHrates>
</ionChannelHH>
<ionChannelHH id="na" conductance="10 pS">
<gateHHrates id="h" instances="1">
<forwardRate type="HHExpRate" midpoint="-65 mV" rate="0.07 per_ms" scale="-20 mV"/>
<q10Settings type="q10ExpTemp" experimentalTemp="32 degC" q10Factor="3"/>
<reverseRate type="HHSigmoidRate" midpoint="-35 mV" rate="1 per_ms" scale="10 mV"/>
</gateHHrates>
<gateHHrates id="m" instances="3">
<forwardRate type="HHExpLinearRate" midpoint="-40 mV" rate="1 per_ms" scale="10 mV"/>
<q10Settings type="q10ExpTemp" experimen
3. Run Reference
from tvbo.adapters.neuroml import run_lems_example= run_lems_example("LEMS_NML2_Ex17_Tissue.xml" )for name, arr in ref_outputs.items():print (f" { name} : shape= { arr. shape} " )
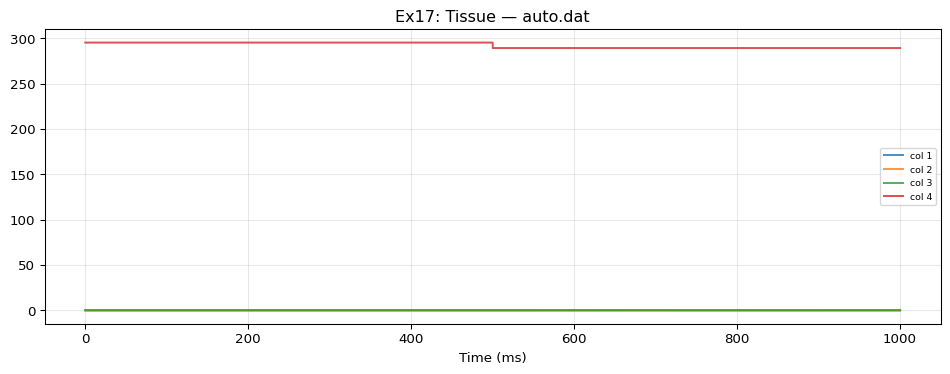
auto.dat: shape=(100001, 5)
4. Run TVBO
= exp.run("neuroml" )= result.integration.dataprint (f"TVBO: { da. dims} , shape= { da. shape} " )
TVBO: ('time', 'variable'), shape=(100001, 1)
5. Compare & Plot
from tvbo.adapters.neuroml import plot_lems_comparison"LEMS_NML2_Ex17_Tissue.xml" , ref_outputs, result.integration.data, title_prefix= "Ex17" )