Model: Instance-Based Network
Three iafCell instances with explicit 3D placement. Cell 0 receives a pulse input and connects to cells 1 and 2 via two different expOneSynapse connections.
Reference: NML2_InstanceBasedNetwork.nml
1. Define Network in TVBO
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NeuroML Ex13: Instance-Based Network" dynamics: name: iaf iri: neuroml:iafCell parameters: leakReversal: { value: -60, unit: mV } thresh: { value: -55, unit: mV } reset: { value: -62, unit: mV } C: { value: 1.0, unit: nF } leakConductance: { value: 0.05, unit: uS } network: number_of_nodes: 4 dynamics: iaf: name: iaf iri: neuroml:iafCell parameters: leakReversal: { value: -60, unit: mV } thresh: { value: -55, unit: mV } reset: { value: -62, unit: mV } C: { value: 1.0, unit: nF } leakConductance: { value: 0.05, unit: uS } pulseGen1: name: pulseGen1 iri: neuroml:pulseGenerator parameters: delay: { value: 100, unit: ms } duration: { value: 100, unit: ms } amplitude: { value: 0.3, unit: nA } nodes: - id: 0 dynamics: iaf - id: 1 dynamics: iaf - id: 2 dynamics: iaf - id: 100 dynamics: pulseGen1 edges: - source: 100 target: 0 - source: 0 target: 1 coupling: expOneSynapse parameters: gbase: { value: 5, unit: nS } erev: { value: 0, unit: mV } tauDecay: { value: 3, unit: ms } - source: 0 target: 2 coupling: expOneSynapse parameters: gbase: { value: 10, unit: nS } erev: { value: 0, unit: mV } tauDecay: { value: 2, unit: ms } integration: method: euler step_size: 0.05 duration: 300.0 time_scale: ms """ )print (f"Model: { exp. dynamics. name if exp. dynamics else 'network' } " )print (f"Network nodes: { len (exp.network.nodes)} " )
Model: iaf
Network nodes: 4
2. Render LEMS XML
= exp.render("lems" )print (xml[:2000 ])
<Lems>
<Target component="sim_NeuroML_Ex13__Instance_Based_Network"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Inputs.xml"/>
<Include file="Simulation.xml"/>
<iafCell id="iaf" C="1 nF" leakConductance="0.05 uS" leakReversal="-60 mV" reset="-62 mV" thresh="-55 mV"/>
<pulseGenerator id="pulseGen1" amplitude="0.3 nA" delay="100.0 ms" duration="100.0 ms"/>
<expOneSynapse id="expOneSynapse" erev="0.0 mV" gbase="5.0 nS" tauDecay="3.0 ms"/>
<expOneSynapse id="expOneSynapse_2" erev="0.0 mV" gbase="10.0 nS" tauDecay="2.0 ms"/>
<network id="net1">
<population id="iaf_pop" component="iaf" size="3"/>
<synapticConnection from="iaf_pop[0]" to="iaf_pop[1]" synapse="expOneSynapse" destination="synapses"/>
<synapticConnection from="iaf_pop[0]" to="iaf_pop[2]" synapse="expOneSynapse_2" destination="synapses"/>
<explicitInput target="iaf_pop[0]" input="pulseGen1" destination="synapses"/>
</network>
<Simulation id="sim_NeuroML_Ex13__Instance_Based_Network" length="300.0ms" step="0.05ms" target="net1">
<OutputFile id="of0" fileName="results/iaf.dat">
<OutputColumn id="iaf_pop_0_v" quantity="iaf_pop[0]/v"/>
<OutputColumn id="iaf_pop_1_v" quantity="iaf_pop[1]/v"/>
<OutputColumn id="iaf_pop_2_v" quantity="iaf_pop[2]/v"/>
</OutputFile>
</Simulation>
</Lems>
3. Run Reference
from tvbo.adapters.neuroml import run_lems_example= run_lems_example("LEMS_NML2_Ex13_Instances.xml" )for name, arr in ref_outputs.items():print (f" { name} : shape= { arr. shape} " )
auto.dat: shape=(6001, 4)
4. Run TVBO
= exp.run("neuroml" )= result.integration.dataprint (f"TVBO: { da. dims} , shape= { da. shape} " )
TVBO: ('time', 'quantity'), shape=(6001, 3)
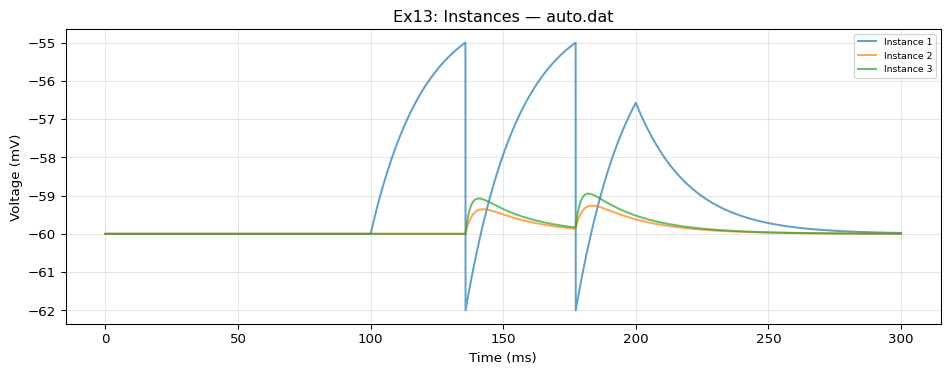
5. Compare & Plot
from tvbo.adapters.neuroml import plot_lems_comparison"LEMS_NML2_Ex13_Instances.xml" , ref_outputs, result.integration.data, title_prefix= "Ex13" )