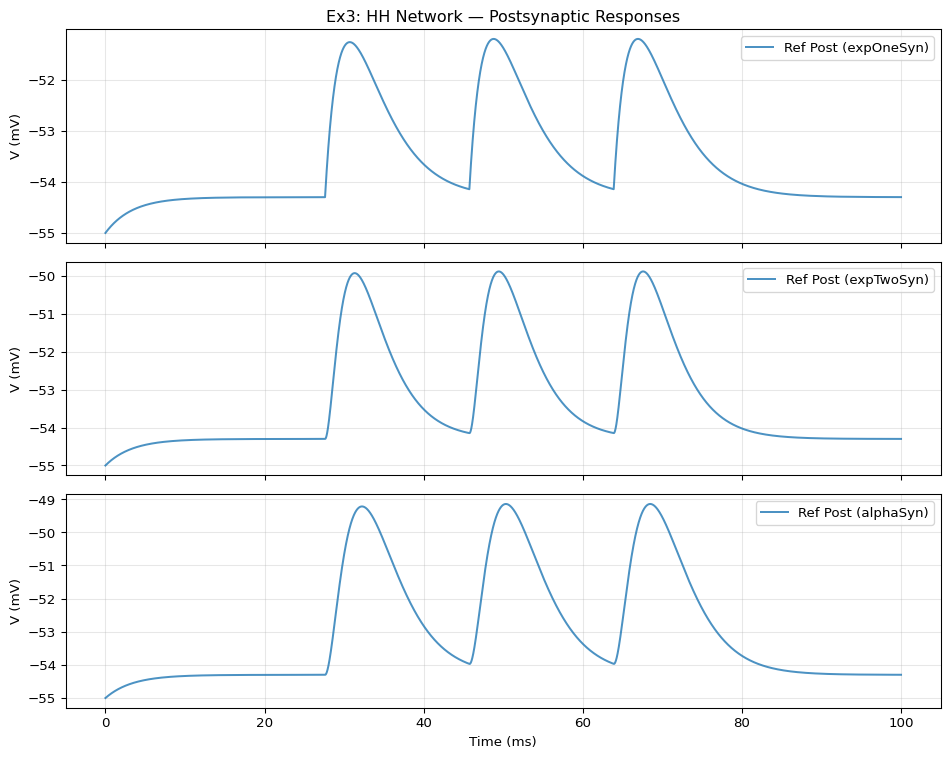
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NeuroML Ex3: HH Network" dynamics: name: hhcell_1 iri: neuroml:pointCellCondBased parameters: C: { value: 10, unit: pF } v0: { value: -65, unit: mV } thresh: { value: 20, unit: mV } components: passive: name: passive iri: neuroml:ionChannelPassive parameters: conductance: { value: 10, unit: pS } number: { value: 300 } erev: { value: -54.3, unit: mV } na: name: na iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 120000 } erev: { value: 50, unit: mV } components: m: name: m iri: neuroml:gateHHrates parameters: { instances: { value: 3 } } components: forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -40, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 4, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -18, unit: mV } h: name: h iri: neuroml:gateHHrates parameters: { instances: { value: 1 } } components: forwardRate: name: forwardRate iri: neuroml:HHExpRate parameters: rate: { value: 0.07, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -20, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHSigmoidRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -35, unit: mV } scale: { value: 10, unit: mV } k: name: k iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 36000 } erev: { value: -77, unit: mV } components: n: name: n iri: neuroml:gateHHrates parameters: { instances: { value: 4 } } components: forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 0.1, unit: per_ms } midpoint: { value: -55, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 0.125, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -80, unit: mV } network: number_of_nodes: 5 dynamics: hhcell_1: name: hhcell_1 iri: neuroml:pointCellCondBased parameters: C: { value: 10, unit: pF } v0: { value: -65, unit: mV } thresh: { value: 20, unit: mV } components: passive: name: passive iri: neuroml:ionChannelPassive parameters: conductance: { value: 10, unit: pS } number: { value: 300 } erev: { value: -54.3, unit: mV } na: name: na iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 120000 } erev: { value: 50, unit: mV } components: m: name: m iri: neuroml:gateHHrates parameters: { instances: { value: 3 } } components: forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -40, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 4, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -18, unit: mV } h: name: h iri: neuroml:gateHHrates parameters: { instances: { value: 1 } } components: forwardRate: name: forwardRate iri: neuroml:HHExpRate parameters: rate: { value: 0.07, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -20, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHSigmoidRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -35, unit: mV } scale: { value: 10, unit: mV } k: name: k iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 36000 } erev: { value: -77, unit: mV } components: n: name: n iri: neuroml:gateHHrates parameters: { instances: { value: 4 } } components: forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 0.1, unit: per_ms } midpoint: { value: -55, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 0.125, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -80, unit: mV } hhcell_2: name: hhcell_2 iri: neuroml:pointCellCondBased parameters: C: { value: 10, unit: pF } v0: { value: -55, unit: mV } thresh: { value: 20, unit: mV } components: leak: name: leak iri: neuroml:ionChannelPassive parameters: conductance: { value: 10, unit: pS } number: { value: 300 } erev: { value: -54.3, unit: mV } nodes: - id: 0 dynamics: hhcell_1 record: false - id: 1 dynamics: hhcell_2 - id: 2 dynamics: hhcell_2 - id: 3 dynamics: hhcell_2 - id: 100 dynamics: pulseGenerator parameters: delay: { value: 25, unit: ms } duration: { value: 50, unit: ms } amplitude: { value: 0.065, unit: nA } edges: - source: 100 target: 0 - source: 0 target: 1 coupling: expOneSynapse parameters: gbase: { value: 0.5, unit: nS } erev: { value: 0, unit: mV } tauDecay: { value: 3, unit: ms } - source: 0 target: 2 coupling: expTwoSynapse parameters: gbase: { value: 0.5, unit: nS } erev: { value: 0, unit: mV } tauRise: { value: 1, unit: ms } tauDecay: { value: 2, unit: ms } - source: 0 target: 3 coupling: alphaSynapse parameters: gbase: { value: 0.5, unit: nS } erev: { value: 0, unit: mV } tau: { value: 2, unit: ms } integration: method: euler step_size: 0.005 duration: 100.0 time_scale: ms """ )print (f"Model: { exp. dynamics. name if exp. dynamics else 'network' } " )print (f"Network nodes: { len (exp.network.nodes)} " )