# =============================================================================
# FitzHugh-Nagumo on a Directed Weighted Brain Network
# =============================================================================
# Source: https://juliadynamics.github.io/NetworkDynamics.jl/stable/generated/directed_and_weighted_graphs/
#
# FitzHugh-Nagumo relaxation oscillators coupled via electrical gap junctions
# on a directed, weighted 90-region brain connectivity (AAL atlas).
# =============================================================================
id: 102
label: "FitzHugh-Nagumo Brain Synchronization"
description: >
Neurodynamic model of synchronization: FitzHugh-Nagumo oscillators
coupled through weighted, directed electrical gap junctions on a
90-region brain atlas (AAL, Tzourio-Mazoyer et al., 2002).
Recreates the NetworkDynamics.jl Directed and Weighted Graphs tutorial.
references:
- "https://juliadynamics.github.io/NetworkDynamics.jl/stable/generated/directed_and_weighted_graphs/"
- "FitzHugh, R. (1961). Impulses and physiological states in theoretical models of nerve membrane. Biophysical Journal, 1(6), 445-466."
- "Nagumo, J., Arimoto, S., & Yoshizawa, S. (1962). An active pulse transmission line simulating nerve axon. Proceedings of the IRE, 50(10), 2061-2070."
- "Tzourio-Mazoyer, N., et al. (2002). Automated anatomical labeling of activations in SPM. Neuroimage, 15(1), 273-289."
# -- Local Dynamics ------------------------------------------------------------
dynamics:
name: FitzHughNagumo
label: "FitzHugh-Nagumo oscillator"
description: >
Two-variable relaxation oscillator. u is the fast excitatory
variable (membrane potential analogue), v is the slow inhibitory
recovery variable.
eps du/dt = u - u^3/3 - v + esum; dv/dt = (u + a) eps
system_type: continuous
autonomous: true
has_reference: "FitzHugh1961"
parameters:
a:
label: "Control parameter"
symbol: "a"
value: 0.5
unit: "1"
description: "Excitability threshold; determines oscillatory vs excitable regime"
epsilon:
label: "Time-scale separation"
symbol: "epsilon"
value: 0.05
unit: "1"
description: "Ratio of fast (u) to slow (v) time scales"
coupling_terms:
c_electrical:
description: "Aggregated electrical gap-junction coupling (sum of incident edge outputs)"
state_variables:
u:
label: "Fast excitatory variable"
symbol: "u"
equation:
rhs: "u - u**3 / 3 - v + c_electrical"
description: "du/dt = u - u^3/3 - v + sum(edges)"
initial_value: 0.0
distribution:
name: Gaussian
seed: 42
domain: { lo: -15.0, hi: 15.0 }
parameters:
mean: { name: mean, value: 0.0 }
std: { name: std, value: 5.0 }
unit: "mV"
variable_of_interest: true
coupling_variable: true
v:
label: "Slow inhibitory variable"
symbol: "v"
equation:
rhs: "(u - a) * epsilon"
description: "dv/dt = (u - a) eps"
initial_value: 0.0
distribution:
name: Gaussian
seed: 42
domain: { lo: -15.0, hi: 15.0 }
parameters:
mean: { name: mean, value: 0.0 }
std: { name: std, value: 5.0 }
unit: "mV"
variable_of_interest: true
# -- Network -------------------------------------------------------------------
network:
label: "AAL-90 directed brain network"
description: >
Directed, weighted structural connectivity from diffusion tensor
imaging. 90 cortical regions from the Automated Anatomical Labeling
(AAL) atlas (Tzourio-Mazoyer et al., 2002). Edge weights are set
at runtime from the Norm_G_DTI.txt matrix.
number_of_nodes: 90
edge_matrix_files:
- ../Norm_G_DTI.txt
parcellation:
label: "AAL-90"
atlas:
name: "AAL"
distance_unit: "mm"
coupling:
ElectricalCoupling:
name: ElectricalCoupling
label: "Weighted electrical gap-junction coupling"
description: >
Directed coupling: each edge computes w * sigma * (u_src - u_dst),
where w is the per-edge structural weight and sigma is a global
coupling strength parameter.
delayed: false
parameters:
w:
label: "Edge weight"
symbol: "w"
value: 1.0
unit: "1"
description: "Structural connectivity weight (per-edge, from DTI)"
sigma:
label: "Global coupling strength"
symbol: "sigma"
value: 0.5
unit: "1"
description: "Global scaling of electrical gap-junction coupling"
pre_expression:
rhs: "w * sigma * (x_j - x_i)"
description: "Directed weighted diffusive coupling"
# -- Integration ---------------------------------------------------------------
integration:
description: >
Automatic stiffness-detecting solver: Tsit5 with fallback to TRBDF2.
dt=0.1 over t in [0, 200].
method: AutoTsit5
step_size: 0.1
duration: 200.0
time_scale: "ms"
FitzHugh-Nagumo Brain Synchronization
Directed weighted coupling on a 90-region brain atlas
Replicates the NetworkDynamics.jl Directed and Weighted Graphs tutorial — FitzHugh-Nagumo oscillators on a 90-region AAL brain connectivity derived from DTI (Tzourio-Mazoyer et al., 2002). The structural connectivity matrix Norm_G_DTI.txt provides directed, weighted edges.
YAML Specification
Model Report
FitzHugh-Nagumo Brain Synchronization
Neurodynamic model of synchronization: FitzHugh-Nagumo oscillators coupled through weighted, directed electrical gap junctions on a 90-region brain atlas (AAL, Tzourio-Mazoyer et al., 2002). Recreates the NetworkDynamics.jl Directed and Weighted Graphs tutorial.
1. Brain Network: AAL-90 directed brain network
Directed, weighted structural connectivity from diffusion tensor imaging. 90 cortical regions from the Automated Anatomical Labeling (AAL) atlas (Tzourio-Mazoyer et al., 2002). Edge weights are set at runtime from the Norm_G_DTI.txt matrix.
- Parcellation: AAL-90 (AAL)
- Regions: 90
Coupling: Weighted electrical gap-junction coupling
Directed coupling: each edge computes w * sigma * (u_src - u_dst), where w is the per-edge structural weight and sigma is a global coupling strength parameter.
Receives \(u\) from connected regions.
Pre-synaptic: \(c_{\text{pre}} = \sigma \cdot w \cdot \left(x_{j} - x_{i}\right)\)
| Parameter | Value | Description |
|---|---|---|
| \(\sigma\) | 0.5 | Global scaling of electrical gap-junction coupling |
| \(w\) | 1.0 | Structural connectivity weight (per-edge, from DTI) |
2. Local Dynamics: FitzHugh-Nagumo oscillator
Two-variable relaxation oscillator. The model comprises 2 state variables.
2.1 State Equations
\[\dot{u} = c_{\mathcal{electri}} + u - v - \frac{u^{3}}{3}\]
\[\dot{v} = \epsilon \cdot \left(u - a\right)\]
2.2 Parameters
| Parameter | Value | Unit | Description |
|---|---|---|---|
| \(a\) | 0.5 | dimensionless | Excitability threshold; determines oscillatory vs excitable regime |
| \(\epsilon\) | 0.05 | dimensionless | Ratio of fast (u) to slow (v) time scales |
3. Numerical Integration
- Method: AutoTsit5
- Time step: \(\Delta t = 0.1\) ms
- Duration: 200.0 ms
References
- https://juliadynamics.github.io/NetworkDynamics.jl/stable/generated/directed_and_weighted_graphs/
- FitzHugh, R. (1961). Impulses and physiological states in theoretical models of nerve membrane. Biophysical Journal, 1(6), 445-466.
- Nagumo, J., Arimoto, S., & Yoshizawa, S. (1962). An active pulse transmission line simulating nerve axon. Proceedings of the IRE, 50(10), 2061-2070.
- Tzourio-Mazoyer, N., et al. (2002). Automated anatomical labeling of activations in SPM. Neuroimage, 15(1), 273-289.
Generated Julia Code
Code(exp.render_code("networkdynamics"), language='julia')using Graphs
using NetworkDynamics
using OrdinaryDiffEqTsit5
using OrdinaryDiffEqSDIRK
using DelimitedFiles
using SimpleWeightedGraphs
function FitzHughNagumo_f!(dx, x, esum, (a, epsilon,), t)
u, v = x
c_electrical = esum[1]
dx[1] = c_electrical + u - v - u .^ 3 / 3
dx[2] = epsilon .* (u - a)
nothing
end
vertex_FitzHughNagumo = VertexModel(;
f = FitzHughNagumo_f!,
g = StateMask(1:1),
sym = [:u, :v],
psym = [:a => 0.5, :epsilon => 0.05],
name = :FitzHughNagumo,
)
function ElectricalCoupling_edge_g!(e_dst, v_src, v_dst, (sigma, w,), t)
e_dst[1] = sigma .* w .* (v_src[1] - v_dst[1])
nothing
end
edge_ElectricalCoupling = EdgeModel(;
g = Directed(ElectricalCoupling_edge_g!),
outsym = [:coupling],
psym = [:sigma => 0.5, :w => 1.0],
name = :ElectricalCoupling,
)
G = readdlm("../Norm_G_DTI.txt", ',', Float64, '\n')
g_weighted = SimpleWeightedDiGraph(G)
edge_weights = getfield.(collect(edges(g_weighted)), :weight)
g = SimpleDiGraph(g_weighted)
nw = Network(g, vertex_FitzHughNagumo, edge_ElectricalCoupling)
p = NWParameter(nw)
p.e[1:ne(g), :w] = edge_weights
s = NWState(nw)
using Random
rng = MersenneTwister(42)
for node in 1:nv(g)
s.v[node, :u] = 0.0 .+ 5.0 .* randn(rng)
end
for node in 1:nv(g)
s.v[node, :v] = 0.0 .+ 5.0 .* randn(rng)
end
tspan = (0.0, 200.0)
prob = ODEProblem(nw, uflat(s), tspan, pflat(p))
sol = solve(prob, AutoTsit5(TRBDF2()); saveat=0.1)
adj_matrix = Float64.(adjacency_matrix(g))
using Random: MersenneTwister
function spring_layout(g; seed=42, iterations=50, k=1.0)
rng = MersenneTwister(seed)
n = nv(g)
pos = randn(rng, n, 2)
for _ in 1:iterations
disp = zeros(n, 2)
for i in 1:n, j in (i+1):n
d = pos[i, :] - pos[j, :]
dist = max(norm(d), 0.01)
rep = k^2 / dist
disp[i, :] .+= d / dist * rep
disp[j, :] .-= d / dist * rep
end
for e in edges(g)
i, j = src(e), dst(e)
d = pos[j, :] - pos[i, :]
dist = max(norm(d), 0.01)
att = dist^2 / k
disp[i, :] .+= d / dist * att
disp[j, :] .-= d / dist * att
end
for i in 1:n
dl = max(norm(disp[i, :]), 0.01)
pos[i, :] .+= disp[i, :] / dl * min(dl, 0.1)
end
end
return pos
end
using LinearAlgebra: norm
node_positions = spring_layout(g)
using Plots
plot(sol; ylabel="state", xlabel="time", title="FitzHughNagumo on 90-node network")
Run & Plot
ts = exp.run(format="networkdynamics")
ts.plot()Detected IPython. Loading juliacall extension. See https://juliapy.github.io/PythonCall.jl/stable/compat/#IPython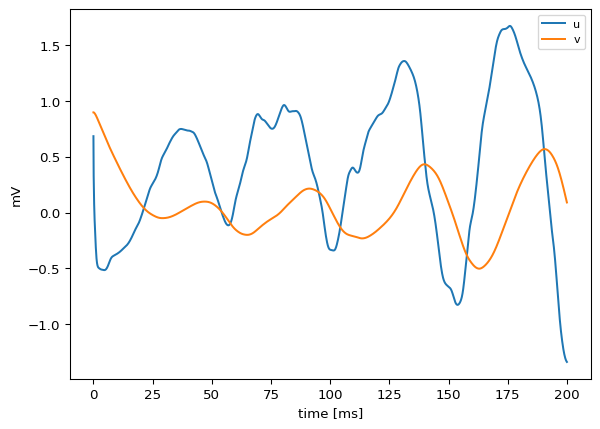
Results
Shape: (2001, 2, 90)
Time range: 0.0 – 200.0
u range: [-16.336, 12.427]
Final u std across nodes: 0.8682
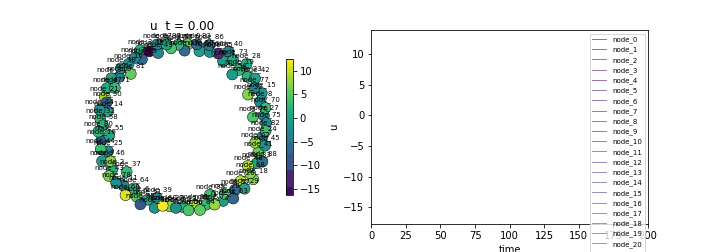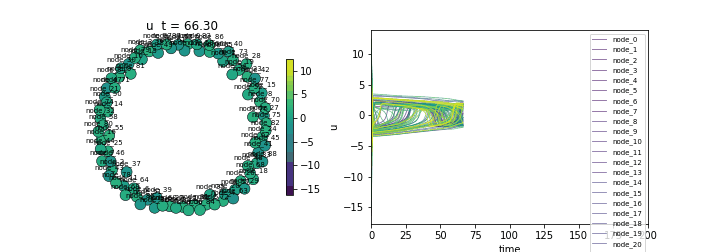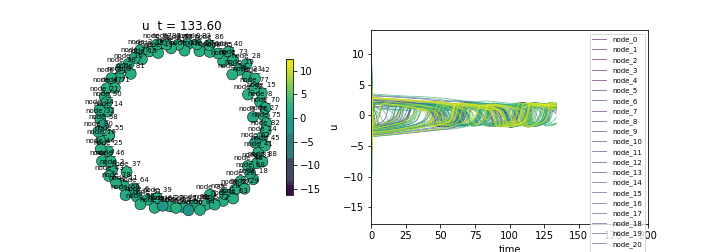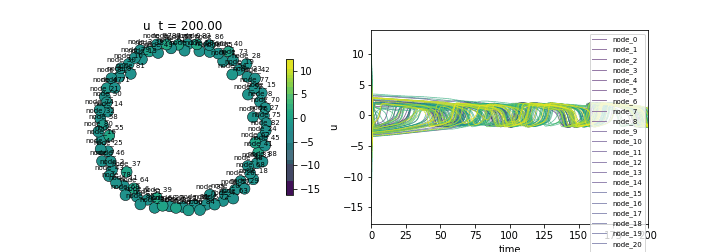
(Lower std indicates synchronization)Animate
ani = ts.animate("u", format="dots")
ani.save("fitzhugh_nagumo.gif", writer="pillow", fps=20)/Users/leonmartin_bih/tools/tvbo/tvbo/plot/animate.py:216: UserWarning: Tight layout not applied. The bottom and top margins cannot be made large enough to accommodate all Axes decorations.
fig.tight_layout()