from tvbo import SimulationExperiment
exp = SimulationExperiment.from_string("""
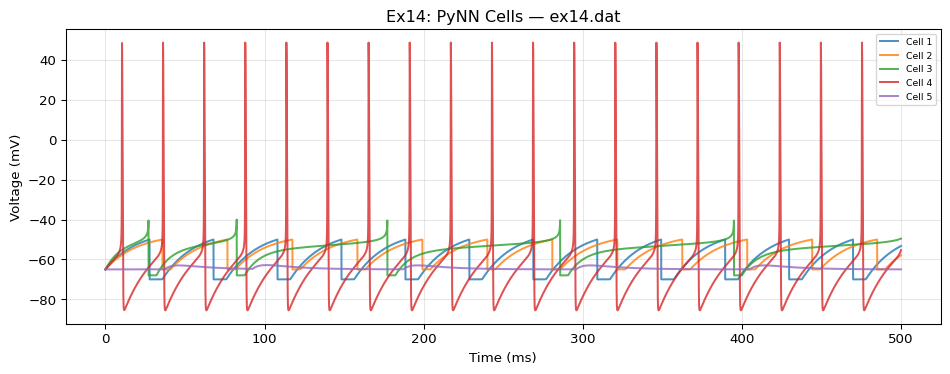
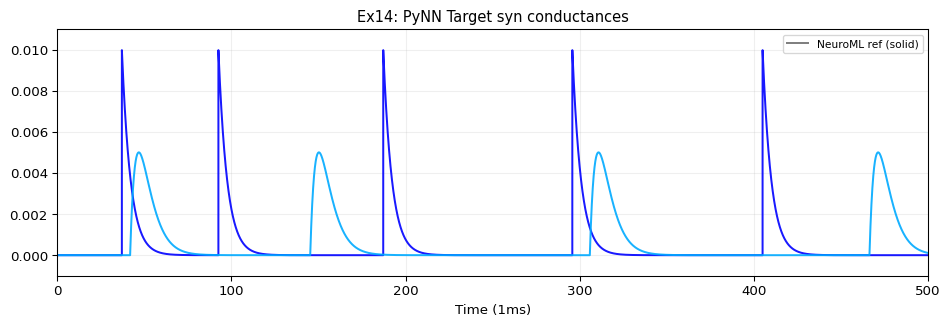
label: "NeuroML Ex14: PyNN Cells"
dynamics:
name: IF_curr_alpha
iri: neuroml:IF_curr_alpha
network:
dynamics:
IF_curr_alpha:
name: IF_curr_alpha
iri: neuroml:IF_curr_alpha
parameters:
cm: {value: 1.0}
i_offset: {value: 0.9}
tau_m: {value: 20.0}
tau_refrac: {value: 10.0}
tau_syn_E: {value: 0.5}
tau_syn_I: {value: 0.5}
v_init: {value: -65}
v_reset: {value: -62.0}
v_rest: {value: -65.0}
v_thresh: {value: -52.0}
IF_curr_exp:
name: IF_curr_exp
iri: neuroml:IF_curr_exp
parameters:
cm: {value: 1.0}
i_offset: {value: 1.0}
tau_m: {value: 20.0}
tau_refrac: {value: 8.0}
tau_syn_E: {value: 5.0}
tau_syn_I: {value: 5.0}
v_init: {value: -65}
v_reset: {value: -70.0}
v_rest: {value: -65.0}
v_thresh: {value: -50.0}
IF_cond_alpha:
name: IF_cond_alpha
iri: neuroml:IF_cond_alpha
parameters:
cm: {value: 1.0}
e_rev_E: {value: 0.0}
e_rev_I: {value: -70.0}
i_offset: {value: 0.9}
tau_m: {value: 20.0}
tau_refrac: {value: 5.0}
tau_syn_E: {value: 0.3}
tau_syn_I: {value: 0.5}
v_init: {value: -65}
v_reset: {value: -65.0}
v_rest: {value: -65.0}
v_thresh: {value: -50.0}
silent_cell:
name: silent_cell
iri: neuroml:IF_cond_alpha
parameters:
cm: {value: 1.0}
e_rev_E: {value: 0.0}
e_rev_I: {value: -70.0}
i_offset: {value: 0}
tau_m: {value: 20.0}
tau_refrac: {value: 5.0}
tau_syn_E: {value: 5}
tau_syn_I: {value: 10}
v_init: {value: -65}
v_reset: {value: -65.0}
v_rest: {value: -65.0}
v_thresh: {value: -50.0}
IF_cond_exp:
name: IF_cond_exp
iri: neuroml:IF_cond_exp
parameters:
cm: {value: 1.0}
e_rev_E: {value: 0.0}
e_rev_I: {value: -70.0}
i_offset: {value: 1.0}
tau_m: {value: 20.0}
tau_refrac: {value: 5.0}
tau_syn_E: {value: 5.0}
tau_syn_I: {value: 5.0}
v_init: {value: -65}
v_reset: {value: -68.0}
v_rest: {value: -65.0}
v_thresh: {value: -52.0}
EIF_cond_exp_isfa_ista:
name: EIF_cond_exp_isfa_ista
iri: neuroml:EIF_cond_exp_isfa_ista
parameters:
a: {value: 0.0}
b: {value: 0.0805}
cm: {value: 0.281}
delta_T: {value: 2.0}
e_rev_E: {value: 0.0}
e_rev_I: {value: -80.0}
i_offset: {value: 0.6}
tau_m: {value: 9.3667}
tau_refrac: {value: 5}
tau_syn_E: {value: 5.0}
tau_syn_I: {value: 5.0}
tau_w: {value: 144.0}
v_init: {value: -65}
v_reset: {value: -68.0}
v_rest: {value: -70.6}
v_spike: {value: -40.0}
v_thresh: {value: -52.0}
EIF_cond_alpha_isfa_ista:
name: EIF_cond_alpha_isfa_ista
iri: neuroml:EIF_cond_alpha_isfa_ista
parameters:
a: {value: 0.0}
b: {value: 0.0805}
cm: {value: 0.281}
delta_T: {value: 0}
e_rev_E: {value: 0.0}
e_rev_I: {value: -80.0}
i_offset: {value: 0.6}
tau_m: {value: 9.3667}
tau_refrac: {value: 5}
tau_syn_E: {value: 5.0}
tau_syn_I: {value: 5.0}
tau_w: {value: 144.0}
v_init: {value: -65}
v_reset: {value: -68.0}
v_rest: {value: -70.6}
v_spike: {value: -40.0}
v_thresh: {value: -52.0}
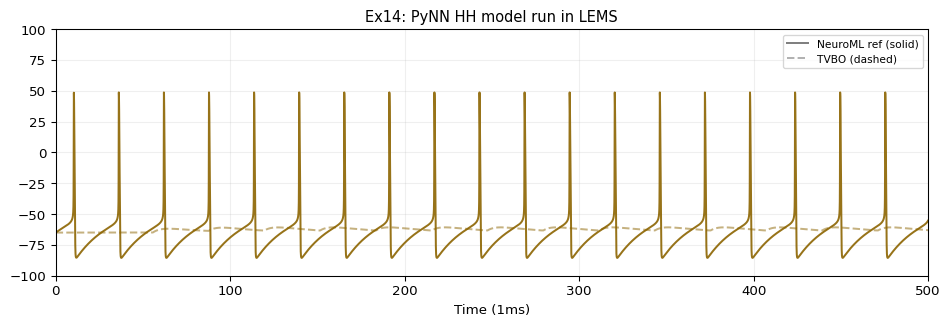
HH_cond_exp:
name: HH_cond_exp
iri: neuroml:HH_cond_exp
parameters:
cm: {value: 0.2}
e_rev_E: {value: 0.0}
e_rev_I: {value: -80.0}
e_rev_K: {value: -90.0}
e_rev_Na: {value: 50.0}
e_rev_leak: {value: -65.0}
g_leak: {value: 0.01}
gbar_K: {value: 6.0}
gbar_Na: {value: 20.0}
i_offset: {value: 0.2}
tau_syn_E: {value: 0.2}
tau_syn_I: {value: 2.0}
v_init: {value: -65}
v_offset: {value: -63.0}
spikes1:
name: spikes1
iri: neuroml:SpikeSourcePoisson
parameters:
start: {value: 50, unit: ms}
duration: {value: 400, unit: ms}
rate: {value: 50, unit: Hz}
spikes2:
name: spikes2
iri: neuroml:SpikeSourcePoisson
parameters:
start: {value: 50, unit: ms}
duration: {value: 300, unit: ms}
rate: {value: 80, unit: Hz}
syn1:
name: syn1
iri: neuroml:expCondSynapse
parameters:
tau_syn: {value: 5}
e_rev: {value: 0}
syn2:
name: syn2
iri: neuroml:alphaCondSynapse
parameters:
tau_syn: {value: 5}
e_rev: {value: 0}
syn3:
name: syn3
iri: neuroml:expCurrSynapse
parameters:
tau_syn: {value: 5}
syn4:
name: syn4
iri: neuroml:alphaCurrSynapse
parameters:
tau_syn: {value: 5}
nodes:
# 7 cell populations (1 each) — only 4 are recorded in reference output
- {id: 0, dynamics: IF_curr_alpha, record: false}
- {id: 1, dynamics: IF_curr_exp}
- {id: 2, dynamics: IF_cond_alpha}
- {id: 3, dynamics: IF_cond_exp, record: false}
- {id: 4, dynamics: EIF_cond_exp_isfa_ista}
- {id: 5, dynamics: EIF_cond_alpha_isfa_ista, record: false}
- {id: 6, dynamics: HH_cond_exp}
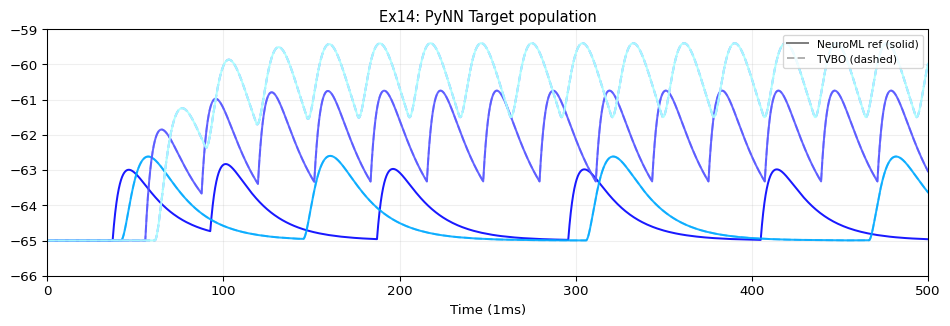
# 4 silent target cells
- {id: 10, dynamics: silent_cell}
- {id: 11, dynamics: silent_cell}
- {id: 12, dynamics: silent_cell}
- {id: 13, dynamics: silent_cell}
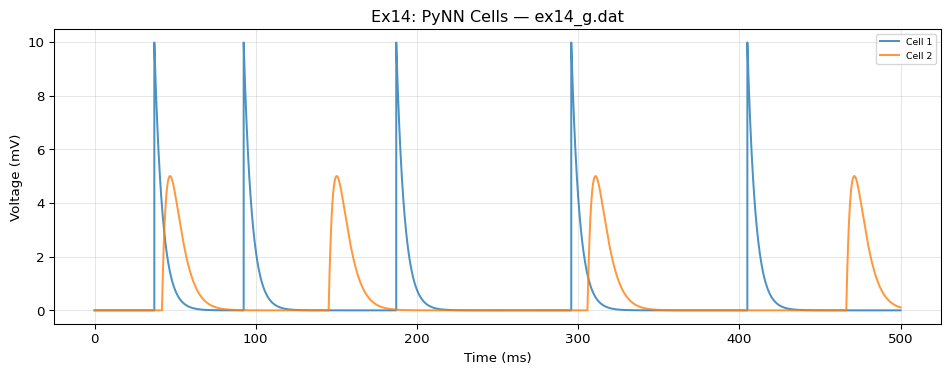
# 2 spike sources (standalone, no connections)
- {id: 20, dynamics: spikes1, record: false}
- {id: 21, dynamics: spikes2, record: false}
edges:
# EIF_cond_exp_isfa_ista → target[0] via expCondSynapse
- {source: 4, target: 10, coupling: syn1, parameters: {weight: {value: 0.01}, delay: {value: 10, unit: ms}}}
# EIF_cond_alpha_isfa_ista → target[1] via alphaCondSynapse
- {source: 5, target: 11, coupling: syn2, parameters: {weight: {value: 0.005}, delay: {value: 20, unit: ms}}}
# IF_curr_alpha → target[2] via expCurrSynapse
- {source: 0, target: 12, coupling: syn3, parameters: {weight: {value: 1}, delay: {value: 30, unit: ms}}}
# IF_cond_exp → target[3] via alphaCurrSynapse
- {source: 3, target: 13, coupling: syn4, parameters: {weight: {value: 0.5}, delay: {value: 40, unit: ms}}}
integration:
method: euler
step_size: 0.01
duration: 500.0
time_scale: ms
""")
dyn_name = next(iter(exp.network.dynamics)) if exp.network and exp.network.dynamics else 'network'
print(f"TVBO model: {dyn_name} (source var: exp)")