Model: iafTauCell
The simplest NeuroML2 example: a leaky integrate-and-fire neuron with a single state variable \(v\) , governed by:
\[\frac{dv}{dt} = \frac{E_L - v}{\tau}\]
With spike event: if \(v > \text{thresh}\) , then \(v \leftarrow \text{reset}\) .
Parameters: leakReversal=-50 mV, thresh=-55 mV, reset=-70 mV, tau=30 ms.
Note: since leakReversal > thresh, this cell will never fire. The NeuroML example also includes iafTauRefCell, iafCell, and iafRefCell variants. The TVBO version matches the iafTauCell (tau-based, no refractory period).
1. Define in TVBO
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NeuroML Ex0: Integrate-and-Fire (iafTau)" dynamics: name: IntegrateAndFire parameters: leakReversal: { value: -50.0, unit: mV } tau: { value: 30.0, unit: ms } thresh: { value: -55.0, unit: mV } reset: { value: -70.0, unit: mV } state_variables: v: equation: { rhs: "(leakReversal - v) / tau" } initial_value: -50.0 unit: mV variable_of_interest: true events: spike: condition: { rhs: "v > thresh" } affect: { rhs: "v = reset" } network: number_of_nodes: 1 integration: method: euler step_size: 0.005 duration: 300.0 time_scale: ms """ )print (f"Model: { exp. dynamics. name if exp. dynamics else 'network' } " )print (f"State variables: { list (exp.dynamics.state_variables.keys())} " )print (f"Events: { list (exp.dynamics.events.keys())} " )
Model: IntegrateAndFire
State variables: ['v']
Events: ['spike']
2. Render LEMS XML
= exp.render("lems" )print (xml[:2000 ])
<Lems>
<!-- Tell jLEMS/jNeuroML which component is the simulation entry point. -->
<Target component="sim_NeuroML_Ex0__Integrate_and_Fire__iafTau_"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Simulation.xml"/>
<!-- ════════════════════════════════════════════════════════════════
Dynamics ComponentType & Component instances
════════════════════════════════════════════════════════════════ -->
<!-- ════════════════════════════════════════════════════════════════
ComponentType: IntegrateAndFire
Generated from TVBO Dynamics: IntegrateAndFire
════════════════════════════════════════════════════════════════ -->
<ComponentType name="IntegrateAndFire">
<!-- Parameters -->
<Parameter name="leakReversal" dimension="voltage"/>
<Parameter name="reset" dimension="voltage"/>
<Parameter name="tau" dimension="time"/>
<Parameter name="thresh" dimension="voltage"/>
<!-- Coupling inputs -->
<!-- Initial condition parameters -->
<Parameter name="v_0" dimension="voltage"/>
<!-- Time conversion for derivatives.
When all parameters and state variables carry proper LEMS dimensions,
LEMS handles unit conversion natively (e.g. tau="30 ms" → 0.03 s).
No SEC constant is needed and TimeDerivatives use the RHS directly.
When dimensions are "none" (dimensionless models), / SEC converts
from model time to SI seconds.
all_dimensioned=True needs_sec=False time_scale=ms -->
<!-- Exposures (one per state variable) -->
<Exposure name="v" dimension="voltage"/>
<Dynamics>
<!-- State variables -->
<StateVariable name="v" dimension="voltage" exposure="v"/>
<!-- Derived variables (simple and conditional/piecewise) -->
<!-- ── Flat dynamics (no spike events) ── -->
<!-- Time derivatives -->
<TimeDerivative variable="v" value="(leakReversal - v)/tau"/>
<!-- Initial condit
3. Run Reference (NeuroML2 via jNeuroML)
from tvbo.adapters.neuroml import run_lems_example, LEMS_EXAMPLES= run_lems_example("LEMS_NML2_Ex0_IaF.xml" )print (f"Reference output files: { list (ref_outputs.keys())} " )for name, arr in ref_outputs.items():print (f" { name} : shape= { arr. shape} " )
Reference output files: ['iaf_v.dat']
iaf_v.dat: shape=(60001, 5)
4. Run TVBO Version
= exp.run("neuroml" )= result.integration.dataprint (f"TVBO: { da. dims} , shape= { da. shape} " )
TVBO: ('time', 'variable'), shape=(60001, 1)
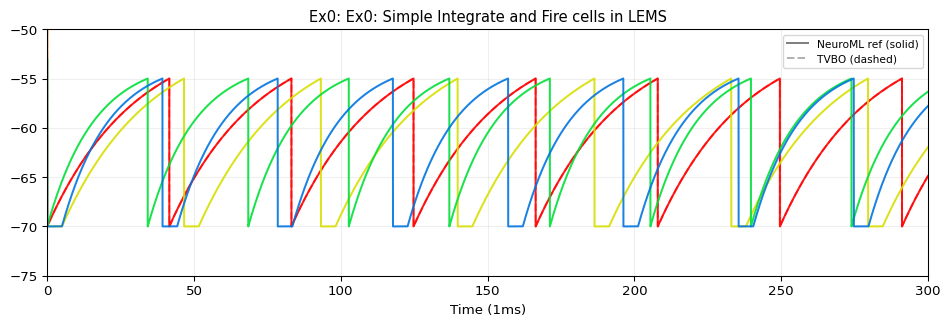
5. Compare & Plot
from tvbo.adapters.neuroml import plot_lems_comparison"LEMS_NML2_Ex0_IaF.xml" , ref_outputs, result.integration.data, title_prefix= "Ex0" )