NML2_AbstractCells.nml
This is the master catalog of all abstract (non-morphological) cell types in NeuroML2. Each cell type is a single-compartment model with analytically defined dynamics.
iafTauCellIntegrateAndFire (Ex0)
1 (v)
iafTauRefCellIaF + refractory
2 (v, lastSpikeTime)
iafCellIaF (conductance-based)
1 (v)
iafRefCellIaF + refractory
2
izhikevichCellIzhikevichCell (Ex2)
2 (v, U)
izhikevich2007CellIzhikevich2007
2 (v, u)
adExIaFCellAdaptiveExponentialIF (Ex8)
2 (v, w)
fitzHughNagumoCellFitzHughNagumo (Ex9)
2 (V, W)
fitzHughNagumo1969CellFHN1969
2 (V, W)
pinskyRinzelCA3CellPinskyRinzel (Ex22)
10
Show Original NML File Structure
import urllib.request= "https://raw.githubusercontent.com/NeuroML/NeuroML2/master/examples/NML2_AbstractCells.nml" with urllib.request.urlopen(url) as resp:= resp.read().decode()# Show component declarations for line in text.split(' \n ' ):= line.strip()if stripped.startswith('<iaf' ) or stripped.startswith('<izh' ) or \ '<adEx' ) or stripped.startswith('<fitz' ) or \ '<pinsky' ):print (stripped[:120 ])
<iafTauCell id="iafTau" leakReversal="-50mV" thresh="-55mV"
<iafTauRefCell id="iafTauRef" leakReversal="-50mV" thresh="-55mV"
<iafCell id="iaf" leakReversal="-50mV" thresh="-55mV"
<iafRefCell id="iafRef" leakReversal="-50mV" thresh="-55mV"
<izhikevichCell id="izBurst" v0 = "-70mV" thresh = "30mV" a="0.02"
<izhikevich2007Cell id="iz2007RS" v0 = "-60mV" C="100 pF" k = "0.7 nS_per_mV"
<adExIaFCell id="adExBurst" C="281pF" gL="30nS" EL="-70.6mV"
<fitzHughNagumoCell id="fn1" I="0.8" />
<fitzHughNagumo1969Cell id="fn1969" a="0.7" b="0.08" I="1.0" phi="0.08" V0="0.0" W0="0.0"/>
<pinskyRinzelCA3Cell id="pr2A" iSoma="0.75 uA_per_cm2" iDend="0 uA_per_cm2" gc="2.1 mS_per_cm2" qd0="0"
Define All Cells in TVBO
FitzHugh-Nagumo 1969
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NML2 AbstractCells: FitzHugh-Nagumo 1969" dynamics: name: FitzHughNagumo1969 parameters: a: { value: 0.7 } b: { value: 0.08 } I: { value: 1.0 } phi: { value: 0.08 } state_variables: V: equation: { rhs: "V - V**3/3 - W + I" } initial_value: 0.0 variable_of_interest: true W: equation: { rhs: "phi*(V + a - b*W)" } initial_value: 0.0 network: number_of_nodes: 1 integration: method: euler step_size: 0.01 duration: 200.0 time_scale: ms """ )print (f"FHN1969 model defined: { exp_fn1969. dynamics. name} " )
FHN1969 model defined: FitzHughNagumo1969
Izhikevich 2007
= SimulationExperiment.from_string(""" label: "NML2 AbstractCells: Izhikevich 2007" dynamics: name: Izhikevich2007 description: "Izhikevich 2007 model" parameters: C: { value: 100.0, description: "Capacitance (pF)" } k: { value: 0.7, description: "nS/mV" } vr: { value: -60.0, description: "Resting potential (mV)" } vt: { value: -40.0, description: "Instantaneous threshold (mV)" } vpeak: { value: 35.0, description: "Spike cutoff (mV)" } a: { value: 0.03, description: "Recovery rate (1/ms)" } b: { value: -2.0, description: "Recovery sensitivity (nS)" } c: { value: -50.0, description: "Reset potential (mV)" } d: { value: 100.0, description: "Recovery reset (pA)" } state_variables: v: equation: { rhs: "(k*(v - vr)*(v - vt) - u) / C" } initial_value: -60.0 variable_of_interest: true u: equation: { rhs: "a*(b*(v - vr) - u)" } initial_value: 0.0 events: spike: condition: { rhs: "v > vpeak" } affect: { rhs: "v = c; u = u + d" } network: number_of_nodes: 1 integration: method: euler step_size: 0.005 duration: 200.0 time_scale: ms """ )print (f"Iz2007 model defined: { exp_iz2007. dynamics. name} " )
Iz2007 model defined: Izhikevich2007
Render and Inspect LEMS
# Show FHN1969 LEMS = exp_fn1969.render("lems" )print ("=== FHN1969 LEMS (excerpt) ===" )print (xml[:1200 ])
=== FHN1969 LEMS (excerpt) ===
<Lems>
<!-- Tell jLEMS/jNeuroML which component is the simulation entry point. -->
<Target component="sim_NML2_AbstractCells__FitzHugh_Nagumo_1969"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Simulation.xml"/>
<!-- ════════════════════════════════════════════════════════════════
Dynamics ComponentType & Component instances
════════════════════════════════════════════════════════════════ -->
<!-- ════════════════════════════════════════════════════════════════
ComponentType: FitzHughNagumo1969
Generated from TVBO Dynamics: FitzHughNagumo1969
════════════════════════════════════════════════════════════════ -->
<ComponentType name="FitzHughNagumo1969">
<!-- Parameters -->
<Parameter name="I" dimension="none"/>
<Parameter name="a" dimension="none"/>
<Parameter name="b" dimension="none"/>
<Parameter name="phi" dimension="none"/>
<!-- Coupling inputs -->
<!-- Initial condition parameters -->
<Parameter name="V_0" dimension="none"/>
<Parameter name="W_0" dimension="none"/>
<!-- Time conversion for derivatives.
When all parameters and state variables ca
Run TVBO
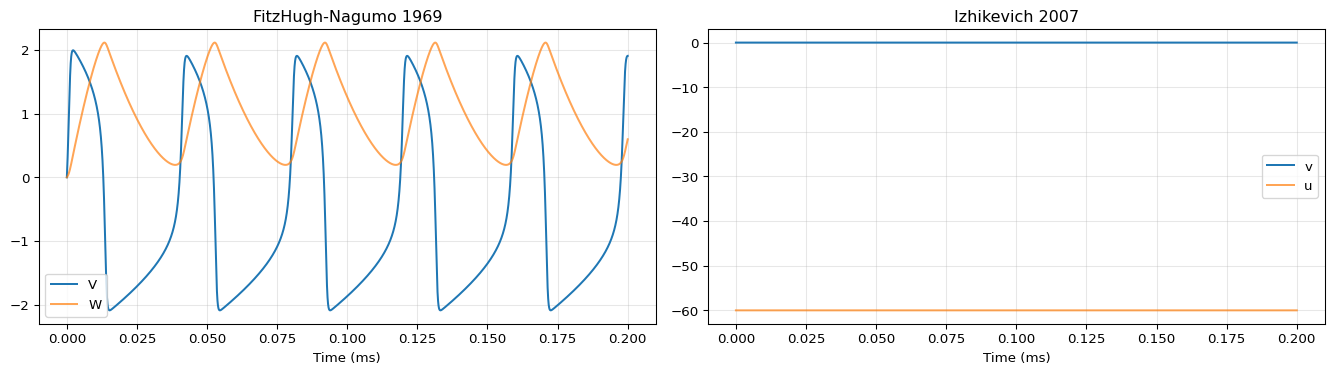
import numpy as npimport matplotlib.pyplot as plt# Run FHN1969 = exp_fn1969.run("neuroml" )= result_fn.integration.data# Run Iz2007 = exp_iz2007.run("neuroml" )= result_iz.integration.data= plt.subplots(1 , 2 , figsize= (14 , 4 ))# FHN1969 = da_fn.coords['time' ].values= da_fn.values0 ].plot(t, vals[:, 0 ], label= 'V' )0 ].plot(t, vals[:, 1 ], label= 'W' , alpha= 0.7 )0 ].set_xlabel("Time (ms)" )0 ].set_title("FitzHugh-Nagumo 1969" )0 ].legend()0 ].grid(True , alpha= 0.3 )# Iz2007 = da_iz.coords['time' ].values= da_iz.values1 ].plot(t, vals[:, 0 ], label= 'v' )1 ].plot(t, vals[:, 1 ], label= 'u' , alpha= 0.7 )1 ].set_xlabel("Time (ms)" )1 ].set_title("Izhikevich 2007" )1 ].legend()1 ].grid(True , alpha= 0.3 )