NML2_AnalogSynapses.nml
Demonstrates graded synaptic transmission using <gradedSynapse> — the postsynaptic current is a continuous function of presynaptic voltage, without discrete spike events. Uses FitzHugh-Nagumo cells.
import urllib.request= "https://raw.githubusercontent.com/NeuroML/NeuroML2/master/examples/NML2_AnalogSynapses.nml" with urllib.request.urlopen(url) as resp:= resp.read().decode()for line in text.split(' \n ' ):= line.strip()if any (tag in stripped for tag in ['<gradedSynapse' , '<silentSynapse' ,'<fitzHugh' , '<network' , '<population' , '<continuousProjection' ,'<continuousConnection' ]):print (stripped[:120 ])
<silentSynapse id="silent1"/>
<silentSynapse id="silent2"/>
<gradedSynapse id="gs2" conductance="5pS" delta="5mV" Vth="-55mV" k="0.025per_ms" erev="0mV"/>
<network id="net1">
<population id="iafPop1" component="iaf" size="1" />
<population id="iafPop2" component="iaf" size="1" />
<population id="iafPop3" component="iaf" size="1" />
<continuousProjection id ="testLinearGradedConn" presynapticPopulation="iafPop1" postsynapticPopulation="iafPop2">
<continuousConnection id="0" preCell="0" postCell="0" preComponent="silent1" postComponent="gs1"/>
<continuousProjection id ="testGradedConn" presynapticPopulation="iafPop1" postsynapticPopulation="iafPop3">
<continuousConnection id="0" preCell="0" postCell="0" preComponent="silent2" postComponent="gs2"/>
TVBO Representation: FHN Cell
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NML2 AnalogSynapses: FHN Cell" dynamics: name: FitzHughNagumo parameters: I: { value: 0.8 } state_variables: V: equation: { rhs: "V - V**3/3 - W + I" } initial_value: 0.0 variable_of_interest: true W: equation: { rhs: "0.08*(V + 0.7 - 0.8*W)" } initial_value: 0.0 network: number_of_nodes: 1 integration: method: euler step_size: 0.01 duration: 200.0 time_scale: s """ )= exp.render("lems" )print (xml[:800 ])
<Lems>
<!-- Tell jLEMS/jNeuroML which component is the simulation entry point. -->
<Target component="sim_NML2_AnalogSynapses__FHN_Cell"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Simulation.xml"/>
<!-- ════════════════════════════════════════════════════════════════
Dynamics ComponentType & Component instances
════════════════════════════════════════════════════════════════ -->
<!-- ════════════════════════════════════════════════════════════════
ComponentType: FitzHughNagumo
Generated from TVBO Dynamics: FitzHughNagumo
════════════════════════════════════════════════════════════════ -->
<ComponentType name="FitzHughNagumo">
<!-- Parameters -->
<Parameter name="I" dimension="none"/>
<!--
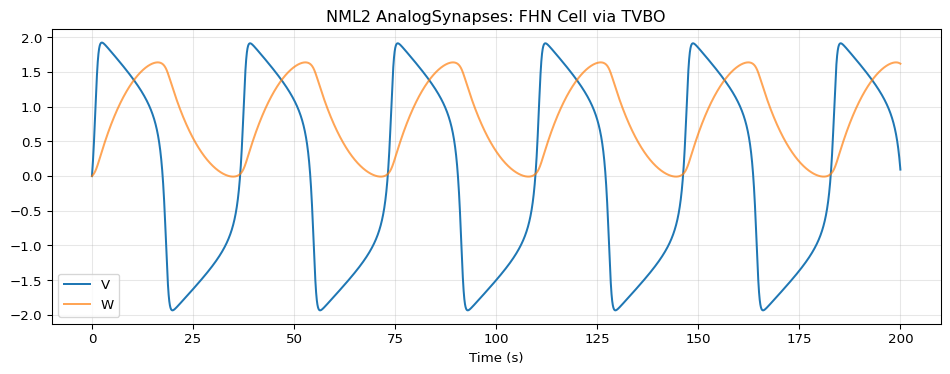
Run TVBO
import numpy as npimport matplotlib.pyplot as plt= exp.run("neuroml" )= result.integration.data= da.coords['time' ].values= da.values= plt.subplots(figsize= (10 , 4 ))0 ], label= 'V' )1 ], label= 'W' , alpha= 0.7 )"Time (s)" )"NML2 AnalogSynapses: FHN Cell via TVBO" )True , alpha= 0.3 )