NML2_GapJunctions.nml
Defines <gapJunction> elements for bidirectional electrical coupling:
<gapJunction id="gj1" conductance="10pS"/>
Connected via <electricalConnection> in the network.
import urllib.request
url = "https://raw.githubusercontent.com/NeuroML/NeuroML2/master/examples/NML2_GapJunctions.nml"
with urllib.request.urlopen(url) as resp:
text = resp.read().decode()
for line in text.split('\n'):
stripped = line.strip()
if '<gapJunction' in stripped or '<electrical' in stripped or \
'<network' in stripped or '<population' in stripped:
print(stripped[:120])
<gapJunction id="gj1" conductance="10pS"/>
<network id="net1">
<population id="iafPop1" component="iaf" size="1" />
<population id="iafPop2" component="iaf" size="1" />
<electricalProjection id ="testGJconn" presynapticPopulation="iafPop1" postsynapticPopulation="iafPop2">
<electricalConnection id="0" preCell="0" postCell="0" synapse="gj1"/>
TVBO Representation: FHN Cell
from tvbo import SimulationExperiment
exp = SimulationExperiment.from_string("""
label: "NML2 GapJunctions: FHN Cell"
dynamics:
name: FitzHughNagumo
parameters:
I: { value: 0.8 }
state_variables:
V:
equation: { rhs: "V - V**3/3 - W + I" }
initial_value: 0.0
variable_of_interest: true
W:
equation: { rhs: "0.08*(V + 0.7 - 0.8*W)" }
initial_value: 0.0
network:
number_of_nodes: 1
integration:
method: euler
step_size: 0.01
duration: 200.0
time_scale: s
""")
xml = exp.render("lems")
print(xml[:800])
<Lems>
<!-- Tell jLEMS/jNeuroML which component is the simulation entry point. -->
<Target component="sim_NML2_GapJunctions__FHN_Cell"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Simulation.xml"/>
<!-- ════════════════════════════════════════════════════════════════
Dynamics ComponentType & Component instances
════════════════════════════════════════════════════════════════ -->
<!-- ════════════════════════════════════════════════════════════════
ComponentType: FitzHughNagumo
Generated from TVBO Dynamics: FitzHughNagumo
════════════════════════════════════════════════════════════════ -->
<ComponentType name="FitzHughNagumo">
<!-- Parameters -->
<Parameter name="I" dimension="none"/>
<!-- Co
Run TVBO
import numpy as np
import matplotlib.pyplot as plt
result = exp.run("neuroml")
da = result.integration.data
t = da.coords['time'].values
vals = da.values
fig, ax = plt.subplots(figsize=(10, 4))
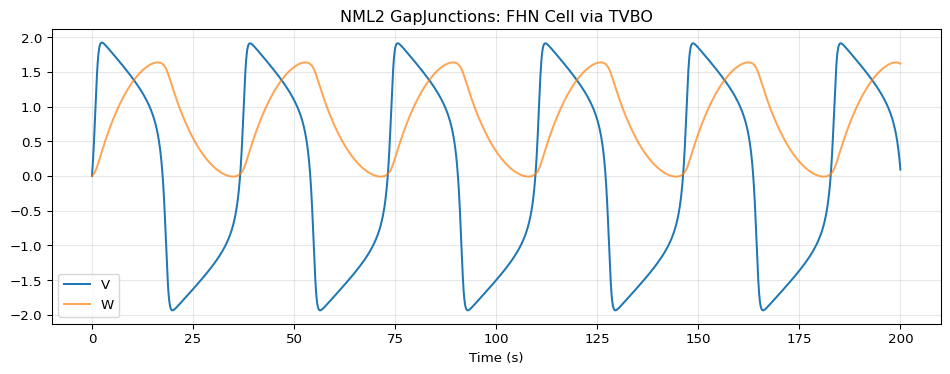
ax.plot(t, vals[:, 0], label='V')
ax.plot(t, vals[:, 1], label='W', alpha=0.7)
ax.set_xlabel("Time (s)")
ax.set_title("NML2 GapJunctions: FHN Cell via TVBO")
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()