NML2_FullNeuroML.nml
A comprehensive example combining all major NeuroML elements in a single file:
Ion channels (Na, K, passive)
Cell with morphology and biophysical properties
Network with populations and projections
Explicit input (pulse generator)
import urllib.request= "https://raw.githubusercontent.com/NeuroML/NeuroML2/master/examples/NML2_FullNeuroML.nml" with urllib.request.urlopen(url) as resp:= resp.read().decode()# Count element types import re= re.findall(r'< ( \w + ) \s ' , text)from collections import Counter= Counter(elements)for el, count in counts.most_common(20 ):print (f" < { el} >: { count} " )
<neuroml>: 1
<morphology>: 1
<segment>: 1
<proximal>: 1
<distal>: 1
<segmentGroup>: 1
<member>: 1
<ionChannelHH>: 1
<expTwoSynapse>: 1
<biophysicalProperties>: 1
<channelDensity>: 1
<spikeThresh>: 1
<specificCapacitance>: 1
<initMembPotential>: 1
<resistivity>: 1
<cell>: 1
<network>: 1
<population>: 1
<projection>: 1
Document Structure
# Show top-level elements for line in text.split(' \n ' ):= line.strip()# Only show opening tags with id= or name=, at first indent levels = len (line) - len (line.lstrip())if indent <= 4 and stripped.startswith('<' ) and not stripped.startswith('<!' ) and \ not stripped.startswith('</' ) and not stripped.startswith('<?' ):print (stripped[:100 ])
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
<morphology id="NeuroMorpho_PyrCell123"> <!-- see FullCell.xml for more details -->
<ionChannelHH id="HH_Na" conductance="10pS" species="na"> <!-- see SimpleIonChannel.xml for more de
<expTwoSynapse id="AMPA" gbase="0.5nS" erev="0mV" tauRise="1ms" tauDecay="2ms" />
<biophysicalProperties id="PyrCellChanDist">
<cell id="PyrCell"
<network id="PyrCellNet">
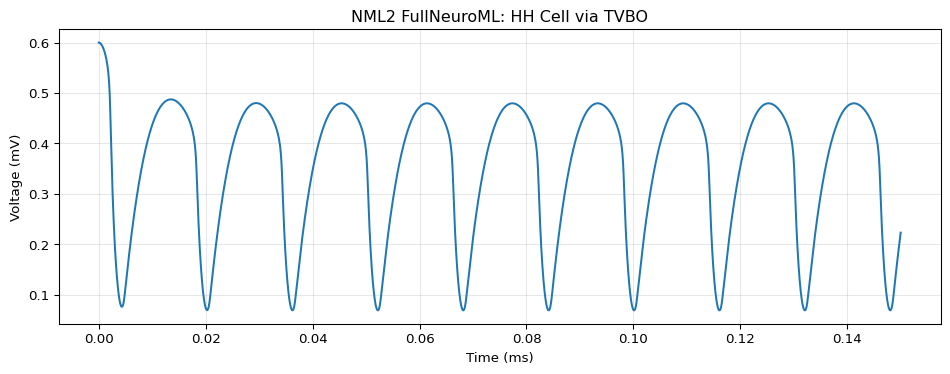
TVBO Representation: HH Cell
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NML2 FullNeuroML: HH" dynamics: name: HodgkinHuxley parameters: C: { value: 10.0 } g_Na: { value: 1200.0 } g_K: { value: 360.0 } g_L: { value: 3.0 } E_Na: { value: 50.0 } E_K: { value: -77.0 } E_L: { value: -54.3 } I_ext: { value: 0.08 } derived_variables: alpha_m: equation: rhs: "Piecewise((1.0, Eq(v, -40.0)), (0.1*(v + 40.0)/(1.0 - exp(-(v + 40.0)/10.0)), True))" beta_m: equation: { rhs: "4.0*exp(-(v + 65.0)/18.0)" } alpha_h: equation: { rhs: "0.07*exp(-(v + 65.0)/20.0)" } beta_h: equation: { rhs: "1.0/(1.0 + exp(-(v + 35.0)/10.0))" } alpha_n: equation: rhs: "Piecewise((0.1, Eq(v, -55.0)), (0.01*(v + 55.0)/(1.0 - exp(-(v + 55.0)/10.0)), True))" beta_n: equation: { rhs: "0.125*exp(-(v + 65.0)/80.0)" } state_variables: v: equation: rhs: "(-g_Na*m**3*h*(v - E_Na) - g_K*n**4*(v - E_K) - g_L*(v - E_L) + I_ext*1000) / C" initial_value: -65.0 variable_of_interest: true m: equation: { rhs: "alpha_m*(1 - m) - beta_m*m" } initial_value: 0.05 h: equation: { rhs: "alpha_h*(1 - h) - beta_h*h" } initial_value: 0.6 n: equation: { rhs: "alpha_n*(1 - n) - beta_n*n" } initial_value: 0.32 network: number_of_nodes: 1 integration: method: euler step_size: 0.01 duration: 150.0 time_scale: ms """ )= exp.render("lems" )print (xml[:800 ])
<Lems>
<!-- Tell jLEMS/jNeuroML which component is the simulation entry point. -->
<Target component="sim_NML2_FullNeuroML__HH"/>
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Simulation.xml"/>
<!-- ════════════════════════════════════════════════════════════════
Dynamics ComponentType & Component instances
════════════════════════════════════════════════════════════════ -->
<!-- ════════════════════════════════════════════════════════════════
ComponentType: HodgkinHuxley
Generated from TVBO Dynamics: HodgkinHuxley
════════════════════════════════════════════════════════════════ -->
<ComponentType name="HodgkinHuxley">
<!-- Parameters -->
<Parameter name="C" dimension="none"/>
<Parameter name="
Run TVBO
import numpy as npimport matplotlib.pyplot as plt= exp.run("neuroml" )= result.integration.data= da.coords['time' ].values= da.values[:, 0 ]= plt.subplots(figsize= (10 , 4 ))"Time (ms)" )"Voltage (mV)" )"NML2 FullNeuroML: HH Cell via TVBO" )True , alpha= 0.3 )