flowchart LR
subgraph TVB["TVB Simulator"]
Sim[Simulator]
Conn[Connectivity]
Mod[Model]
Surf[CorticalSurface]
RM[RegionMapping]
end
subgraph TVBO["TVBO"]
Exp[SimulationExperiment]
Net[Network]
Dyn[Dynamics]
end
Sim -->|from_tvb_simulator| Exp
Conn -->|from_tvb| Net
Surf -->|from_tvb_surface| Net
RM -->|from_tvb_surface| Net
Exp -->|execute / run| Sim
Net -->|"execute(format='tvb')"| Conn
Dyn -->|"execute(format='tvb')"| Mod
style TVB fill:none,stroke:#555
style TVBO fill:none,stroke:#555
The Virtual Brain (TVB)
Bidirectional interoperability with TVB simulator and data types
TVBO provides full bidirectional integration with The Virtual Brain (TVB). You can import TVB objects into tvbo, modify or persist them, and export back to TVB — or use TVB as a simulation backend directly.
Integration Overview
| Direction | Method | Input | Output |
|---|---|---|---|
| TVB → tvbo | Network.from_tvb(conn) |
Connectivity |
Network |
| TVB → tvbo | Network.from_tvb_zip(path) |
ZIP file | Network |
| TVB → tvbo | Network.from_tvb_surface(conn, surf, rmap) |
Conn + Surface + RegionMapping | (Network, Network) |
| TVB → tvbo | SimulationExperiment.from_tvb_simulator(sim) |
Simulator |
SimulationExperiment |
| tvbo → TVB | net.execute(format="tvb") |
Network |
Connectivity |
| tvbo → TVB | dyn.execute(format="tvb") |
Dynamics |
Model |
| tvbo → TVB | exp.execute(format="tvb") |
SimulationExperiment |
Simulator |
| tvbo → TVB | exp.run(format="tvb") |
SimulationExperiment |
TimeSeries |
Pages
| Page | Description |
|---|---|
| Connectome Round-Trip | Lossless connectivity import/export with HDF5 persistence and surface meshes |
Quick Examples
Import a TVB Simulator
Convert a fully configured TVB Simulator into a tvbo SimulationExperiment — preserving the model, coupling, integrator, monitors, noise, and network:
import numpy as np
from tvbo import Dynamics, SimulationExperiment, Network
# Build a TVB simulator the usual way
from tvb.simulator.lab import *
conn = connectivity.Connectivity.from_file()
conn.configure()
conn.speed = np.array([3.0])
sim = simulator.Simulator(
model=models.JansenRit(),
coupling=coupling.SigmoidalJansenRit(
a=np.array([1.0]),
cmax=np.array([0.005]),
midpoint=np.array([6.0]),
r=np.array([0.56]),
),
integrator=integrators.HeunStochastic(
dt=0.1,
noise=noise.Additive(nsig=np.array([0.01])),
),
connectivity=conn,
monitors=[monitors.Raw(), monitors.TemporalAverage(period=1.0)],
simulation_length=1000.0,
)
sim.configure()
print(f"TVB Simulator: {sim.model.__class__.__name__}, "
f"{conn.weights.shape[0]} regions, dt={sim.integrator.dt}")TVB Simulator: JansenRit, 76 regions, dt=0.1exp = SimulationExperiment.from_tvb_simulator(sim)
print(f"Model: {exp.model}")
print(f"Coupling: {exp.coupling.name}")
print(f"Integrator: {exp.integration.method}")
print(f"Duration: {exp.integration.duration} ms")
print(f"dt: {exp.integration.step_size}")
print(f"Noise: {'additive' if exp.integration.noise.additive else 'multiplicative'}")
print(f"Monitors: {list(exp.observations.keys())}")Model: JansenRit
Coupling: SigmoidalJansenRit
Integrator: Heun
Duration: 1000.0 ms
dt: 0.1
Noise: additive
Monitors: ['Raw', 'TemporalAverage']The imported experiment is a standard tvbo object — you can inspect the YAML representation, modify parameters, or re-export:
# Show the YAML specification
print(exp.to_yaml()[:1500])
print("...")id: 244030253939739112725837270727559473241
model: JansenRit
dynamics:
name: JansenRit
parameters:
A:
name: A
value: !!float 'np.float64(3.25)'
description: Maximum amplitude of EPSP [mV]
unit: Millivolt
B:
name: B
value: !!float 'np.float64(22.0)'
description: Maximum amplitude of IPSP [mV]
unit: Millivolt
a:
name: a
value: !!float 'np.float64(0.1)'
description: Reciprocal of the time constant of passive membrane and all other
spatially distributed delays in the dendritic network. Also called average
synaptic time constant.
unit: ms^-1
b:
name: b
value: !!float 'np.float64(0.05)'
description: Rate constant of the inhibitory post-synaptic potential (IPSP)
unit: ms^-1
v0:
name: v0
value: !!float 'np.float64(5.52)'
description: Average firing threshold (PSP) for which half of the firing rate
is achieved
unit: mV
nu_max:
name: nu_max
value: !!float 'np.float64(0.0025)'
description: Asymptotic of the sigmoid function Sigm_JR corresponds to the maximum
firing rate of the neural populations
unit: ms^-1
r:
name: r
value: !!float 'np.float64(0.56)'
description: Steepness (or gain) parameter of the sigmoid function Sigm_JR
unit: mV^-1
J:
name: J
value: !!float 'np.float64(135.0)'
description: Average number of synapses between three ne
...Export a Dynamics Model to TVB
Any tvbo Dynamics can be compiled into a TVB-compatible Model object. This uses Mako code generation to produce a TVB Model subclass:
dyn = Dynamics.from_db("JansenRit")
tvb_model = dyn.execute(format="tvb")
print(f"TVB Model: {tvb_model.__class__.__name__}")
print(f"State vars: {tvb_model.state_variables}")
print(f"Parameters: {tvb_model.parameter_names}")TVB Model: JansenRit
State vars: ['y0', 'y1', 'y2', 'y3', 'y4', 'y5']
Parameters: ['A', 'B', 'J', 'a_1', 'a_2', 'a_3', 'a_4', 'a', 'b', 'mu', 'nu_max', 'r', 'v0']You can inspect the generated TVB model code:
code = dyn.render_code(format="tvb")
# Show first 40 lines
for line in code.splitlines()[:40]:
print(line)
print("...")# Auto-generated standalone model file
import numpy as np
from tvb.basic.neotraits.api import Attr, Final, List, NArray, Range
from tvb.simulator.models.base import Model
class JansenRit(Model):
A = NArray(
label=r":math:`A`",
default=np.array([3.25]),
domain=Range(lo=2.6, hi=9.75, step=0.05),
doc="""Maximum amplitude of EPSP [mV]""",
)
B = NArray(
label=r":math:`B`",
default=np.array([22.0]),
domain=Range(lo=17.6, hi=110.0, step=0.2),
doc="""Maximum amplitude of IPSP [mV]""",
)
J = NArray(
label=r":math:`J`",
default=np.array([135.0]),
domain=Range(lo=65.0, hi=1350.0, step=1.0),
doc="""Average number of synapses between three neuronal populations of the model""",
)
a_1 = NArray(
label=r":math:`a_1`",
default=np.array([1.0]),
domain=Range(lo=0.5, hi=1.5, step=0.1),
doc="""Average probability constant of the number of synapses made by the pyramidal cells to the dendrites of the excitatory interneurons (feedback excitatory loop)""",
)
a_2 = NArray(
label=r":math:`a_2`",
default=np.array([0.8]),
domain=Range(lo=0.4, hi=1.2, step=0.1),
doc="""Average probability constant of the number of synapses made by the EINs to the dendrites of the PCs""",
)
a_3 = NArray(
label=r":math:`a_3`",
...Define and Run a Simulation
A SimulationExperiment can also be defined from a YAML string — useful for sharing reproducible experiment specifications:
# Define an experiment from YAML
exp2 = SimulationExperiment.from_string("""
label: Quick JansenRit demo
model:
name: JansenRit
coupling:
name: SigmoidalJansenRit
parameters:
a: {value: 1.0}
cmax: {value: 0.005}
midpoint: {value: 6.0}
r: {value: 0.56}
integration:
method: HeunStochastic
duration: 500
step_size: 0.1
noise:
additive: true
parameters:
sigma: {value: 0.01}
observations:
Raw:
name: Raw
""")
exp2.network = Network.from_tvb(conn)
print(f"Model: {exp2.model}")
print(f"Method: {exp2.integration.method}")
print(f"Duration: {exp2.integration.duration} ms")
print(f"Step: {exp2.integration.step_size}")Model: JsonObj(name='JansenRit')
Method: HeunStochastic
Duration: 500.0 ms
Step: 0.1Run a simulation using the TVB simulator we already built:
# Run using the previously configured TVB simulator
results = sim.run(simulation_length=500)
(raw_time, raw_data), (tavg_time, tavg_data) = results
import matplotlib.pyplot as plt
fig, axes = plt.subplots(2, 1, figsize=(10, 5), sharex=True)
t = raw_time / 1000 # ms → s
# Plot PSP (y0 - y1) — typical JR output
axes[0].plot(t, raw_data[:, 1, :, 0] - raw_data[:, 2, :, 0],
alpha=0.3, linewidth=0.5)
axes[0].set_ylabel("PSP (y₀ − y₁)")
axes[0].set_title("Jansen-Rit EEG proxy across 76 regions")
# Plot y0 (excitatory PSP)
axes[1].plot(t, raw_data[:, 0, :, 0], alpha=0.3, linewidth=0.5)
axes[1].set_ylabel("y₀")
axes[1].set_xlabel("Time (s)")
plt.tight_layout()
plt.show()/Users/leonmartin_bih/tools/tvbo/.venv/lib/python3.13/site-packages/numba/np/ufunc/gufunc.py:263: RuntimeWarning: overflow encountered in _numba_dfun_jr
return self.ufunc(*args, **kwargs)
/Users/leonmartin_bih/tools/tvbo/.venv/lib/python3.13/site-packages/tvb/simulator/coupling.py:372: RuntimeWarning: overflow encountered in exp
(self.cmax - self.cmin) / (1.0 + numpy.exp(self.r * (self.midpoint - (x_j[:, 0] - x_j[:, 1]))))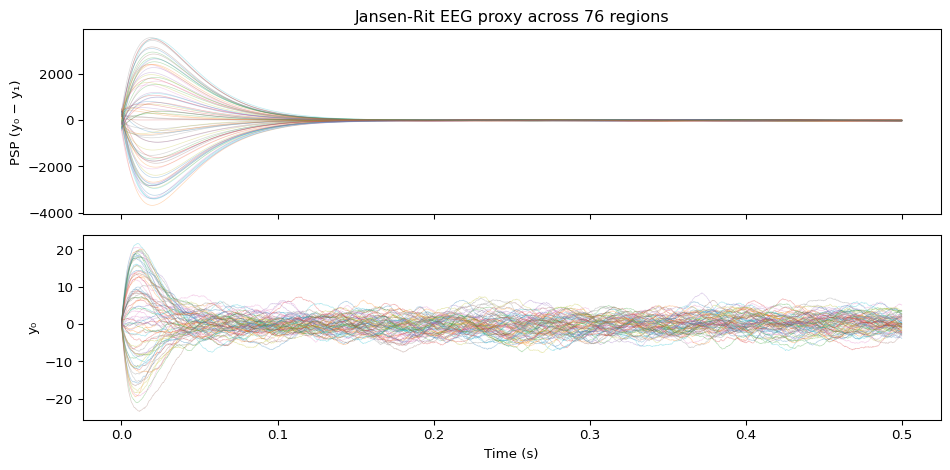
Import a Connectivity
For connectome-only workflows, import TVB connectivity without a full simulator. See the Connectome Round-Trip page for the complete walkthrough including surface meshes and HDF5 persistence.
from tvb.datatypes.connectivity import Connectivity
conn = Connectivity.from_file()
conn.configure()
net = Network.from_tvb(conn)
print(f"{net.number_of_nodes} regions, "
f"weights: [{net.weights_matrix.min():.3f}, {net.weights_matrix.max():.3f}]")76 regions, weights: [0.000, 3.000]net.plot_matrix()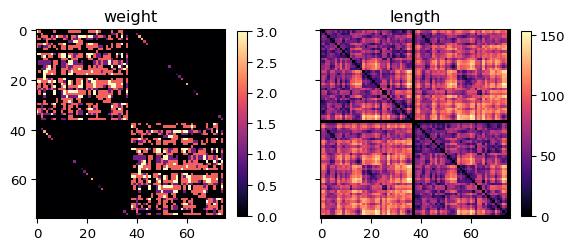
The round-trip is lossless:
import numpy as np
conn2 = net.execute(format="tvb")
np.testing.assert_allclose(conn2.weights, conn.weights)
np.testing.assert_allclose(conn2.tract_lengths, conn.tract_lengths)
print("✓ Weights identical")
print("✓ Tract lengths identical")✓ Weights identical
✓ Tract lengths identical