Model: Hodgkin-Huxley Point Cell
The classic 4-variable conductance-based model:
\[C\frac{dv}{dt} = -g_{Na}\,m^3\,h\,(v - E_{Na}) - g_K\,n^4\,(v - E_K) - g_L\,(v - E_L) + I_{\text{ext}}(t)\]
With gating variable kinetics: \[\frac{dx}{dt} = \alpha_x(v)(1-x) - \beta_x(v)\,x \quad \text{for } x \in \{m, h, n\}\]
The NeuroML2 example uses a pointCellCondBased with separate ion channel populations. By annotating the TVBO dynamics with iri: neuroml:pointCellCondBased and structuring the ion channels as components, TVBO emits standard NeuroML2 types that preserve the jLEMS child-before-parent RK4 evaluation order — giving exact numerical identity with the reference simulation.
1. Define in TVBO
from tvbo import SimulationExperiment# HH parameters matching NeuroML2 Ex1 (hhpointcell) # Uses `iri: neuroml:*` annotations to emit standard NeuroML types. = SimulationExperiment.from_string(""" label: "NeuroML Ex1: Hodgkin-Huxley" dynamics: name: HodgkinHuxley iri: neuroml:pointCellCondBased description: > Hodgkin-Huxley model matching NeuroML2 Ex1. Uses standard NeuroML types via hierarchical components. parameters: C: { value: 10, unit: pF } v0: { value: -65, unit: mV } thresh: { value: 20, unit: mV } pulse_delay: { value: 50, unit: ms } pulse_duration: { value: 50, unit: ms } I_amp: { value: 0.08, unit: nA } components: passive: name: passive iri: neuroml:ionChannelPassive parameters: conductance: { value: 10, unit: pS } number: { value: 300 } erev: { value: -54.3, unit: mV } na: name: na iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 120000 } erev: { value: 50, unit: mV } components: m: name: m iri: neuroml:gateHHrates parameters: instances: { value: 3 } components: forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -40, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 4, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -18, unit: mV } h: name: h iri: neuroml:gateHHrates parameters: instances: { value: 1 } components: forwardRate: name: forwardRate iri: neuroml:HHExpRate parameters: rate: { value: 0.07, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -20, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHSigmoidRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -35, unit: mV } scale: { value: 10, unit: mV } k: name: k iri: neuroml:ionChannelHH parameters: conductance: { value: 10, unit: pS } number: { value: 36000 } erev: { value: -77, unit: mV } components: n: name: n iri: neuroml:gateHHrates parameters: instances: { value: 4 } components: forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 0.1, unit: per_ms } midpoint: { value: -55, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 0.125, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -80, unit: mV } integration: step_size: 0.01 duration: 150.0 time_scale: ms """ )print (f"Model: { exp. dynamics. name if exp. dynamics else 'network' } " )print (f"IRI: { exp. dynamics. iri} " )print (f"Components: { list (exp.dynamics.components.keys())} " )
Model: HodgkinHuxley
IRI: neuroml:pointCellCondBased
Components: ['k', 'na', 'passive']
This example uses iri: neuroml:pointCellCondBased with nested components for ion channels (ionChannelHH, ionChannelPassive) and gates (gateHHrates). TVBO’s adapter detects these annotations and emits standard NeuroML2 XML elements rather than custom ComponentType definitions. This preserves the hierarchical RK4 evaluation order in jLEMS, giving exact numerical identity with the reference simulation at the original step size (0.01 ms) — no gate_rate_scale hack needed.
2. Render LEMS XML
= exp.render("lems" )# Show key elements of the generated XML for line in xml.split(' \n ' ):= line.strip()if any (tag in line_stripped for tag in ['<ionChannel' , '<gateHH' , '<pointCell' ,'<channelPopulation' , '<pulseGenerator' , 'Rate type=' , '<Include' ]):print (line_stripped)
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Inputs.xml"/>
<Include file="Simulation.xml"/>
<ionChannelHH id="k" conductance="10 pS">
<gateHHrates id="n" instances="4">
<forwardRate type="HHExpLinearRate" midpoint="-55 mV" rate="0.1 per_ms" scale="10 mV"/>
<reverseRate type="HHExpRate" midpoint="-65 mV" rate="0.125 per_ms" scale="-80 mV"/>
<ionChannelHH id="na" conductance="10 pS">
<gateHHrates id="h" instances="1">
<forwardRate type="HHExpRate" midpoint="-65 mV" rate="0.07 per_ms" scale="-20 mV"/>
<reverseRate type="HHSigmoidRate" midpoint="-35 mV" rate="1 per_ms" scale="10 mV"/>
<gateHHrates id="m" instances="3">
<forwardRate type="HHExpLinearRate" midpoint="-40 mV" rate="1 per_ms" scale="10 mV"/>
<reverseRate type="HHExpRate" midpoint="-65 mV" rate="4 per_ms" scale="-18 mV"/>
<ionChannelPassive id="passive" conductance="10 pS"/>
<pointCellCondBased id="HodgkinHuxley" C="10 pF" v0="-65 mV" thresh="20 mV">
<channelPopulation id="k_pop" ionChannel="k" number="36000" erev="-77 mV"/>
<channelPopulation id="na_pop" ionChannel="na" number="120000" erev="50 mV"/>
<channelPopulation id="passive_pop" ionChannel="passive" number="300" erev="-54.3 mV"/>
<pulseGenerator id="pulseGen1" delay="50 ms" duration="50 ms" amplitude="0.08 nA"/>
3. Run Reference
from tvbo.adapters.neuroml import run_lems_example= run_lems_example("LEMS_NML2_Ex1_HH.xml" )for name, arr in ref_outputs.items():print (f" { name} : shape= { arr. shape} , t=[ { arr[0 ,0 ]:.4f} , { arr[- 1 ,0 ]:.4f} ]" )
hh_v.dat: shape=(15001, 2), t=[0.0000, 0.1500]
4. Run TVBO Version
= exp.run("neuroml" )= result.integration.dataprint (f"TVBO: { da. dims} , shape= { da. shape} " )
TVBO: ('time', 'variable'), shape=(15001, 1)
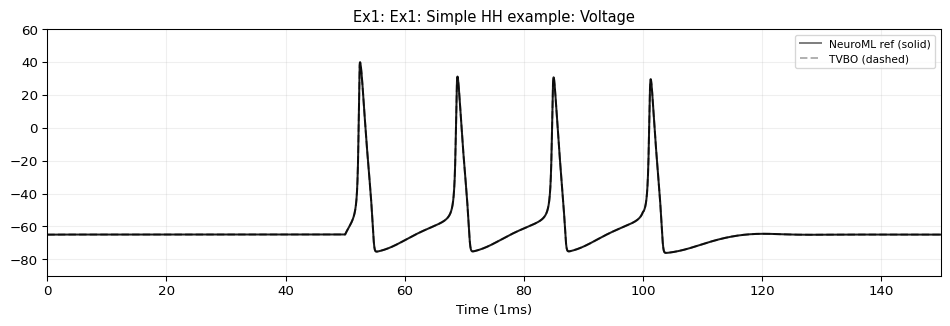
5. Compare & Plot
from tvbo.adapters.neuroml import plot_lems_comparison"LEMS_NML2_Ex1_HH.xml" , ref_outputs, result.integration.data, title_prefix= "Ex1" )
Native Custom YAML (No IRI Dependencies)
The same Hodgkin-Huxley model, defined with full equations — no reference to standard NeuroML component types. Uses iri: extends:base* to declare LEMS base type compatibility while providing all dynamics explicitly. This makes the YAML interoperable with any backend, not just NeuroML.
6. Define Custom Version
= SimulationExperiment.from_string(""" label: "Ex1 HH Custom" dynamics: name: HodgkinHuxleyCustom label: customPointCellCondBased iri: "extends:baseCellMembPot" description: > Hodgkin-Huxley with fully explicit equations. All component types defined from scratch — no standard NeuroML types used. parameters: C: { value: 10, unit: pF } v0: { value: -65, unit: mV } thresh: { value: 20, unit: mV } pulse_delay: { value: 50, unit: ms } pulse_duration: { value: 50, unit: ms } I_amp: { value: 0.08, unit: nA } state_variables: v: equation: { rhs: "(iChannels + iSyn) / C" } initial_value: -65.0 unit: mV variable_of_interest: true derived_variables: iChannels: equation: { rhs: "0" } label: "select:populations[*]/i reduce:add" unit: nA iSyn: equation: { rhs: "0" } label: "select:synapses[*]/i reduce:add required:false" unit: nA events: spike: condition: { rhs: "v > thresh" } components: passive: name: passive label: customIonChannelHH iri: "extends:baseIonChannel" parameters: conductance: { value: 10, unit: pS } number: { value: 300 } erev: { value: -54.3, unit: mV } derived_variables: fopen: equation: { rhs: "1" } g: equation: { rhs: "conductance * fopen" } unit: nS na: name: na label: customIonChannelHH iri: "extends:baseIonChannel" parameters: conductance: { value: 10, unit: pS } number: { value: 120000 } erev: { value: 50, unit: mV } derived_variables: fopen: equation: { rhs: "1" } label: "select:gates[*]/fcond reduce:multiply" g: equation: { rhs: "conductance * fopen" } unit: nS components: m: name: m label: customGateHHrates iri: "extends:baseGate" parameters: instances: { value: 3 } state_variables: q: equation: { rhs: "(inf - q) / tau" } initial_value: 0.05 derived_variables: alpha: equation: { rhs: "0" } label: "select:forwardRate/r" unit: per_ms beta: equation: { rhs: "0" } label: "select:reverseRate/r" unit: per_ms inf: equation: { rhs: "alpha / (alpha + beta)" } tau: equation: { rhs: "1 / (alpha + beta)" } unit: ms fcond: equation: { rhs: "q ** instances" } components: forwardRate: name: forwardRate label: customHHExpLinearRate iri: "extends:baseVoltageDepRate" parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -40, unit: mV } scale: { value: 10, unit: mV } derived_variables: x: equation: { rhs: "(v - midpoint) / scale" } r: conditional: true cases: - condition: "x != 0" equation: { rhs: "rate * x / (1 - exp(-x))" } - condition: "True" equation: { rhs: "rate" } reverseRate: name: reverseRate label: customHHExpRate iri: "extends:baseVoltageDepRate" parameters: rate: { value: 4, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -18, unit: mV } derived_variables: r: equation: { rhs: "rate * exp((v - midpoint) / scale)" } unit: per_ms h: name: h label: customGateHHrates iri: "extends:baseGate" parameters: instances: { value: 1 } state_variables: q: equation: { rhs: "(inf - q) / tau" } initial_value: 0.60 derived_variables: alpha: equation: { rhs: "0" } label: "select:forwardRate/r" unit: per_ms beta: equation: { rhs: "0" } label: "select:reverseRate/r" unit: per_ms inf: equation: { rhs: "alpha / (alpha + beta)" } tau: equation: { rhs: "1 / (alpha + beta)" } unit: ms fcond: equation: { rhs: "q ** instances" } components: forwardRate: name: forwardRate label: customHHExpRate iri: "extends:baseVoltageDepRate" parameters: rate: { value: 0.07, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -20, unit: mV } derived_variables: r: equation: { rhs: "rate * exp((v - midpoint) / scale)" } unit: per_ms reverseRate: name: reverseRate label: customHHSigmoidRate iri: "extends:baseVoltageDepRate" parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -35, unit: mV } scale: { value: 10, unit: mV } derived_variables: r: equation: { rhs: "rate / (1 + exp(-(v - midpoint) / scale))" } unit: per_ms k: name: k label: customIonChannelHH iri: "extends:baseIonChannel" parameters: conductance: { value: 10, unit: pS } number: { value: 36000 } erev: { value: -77, unit: mV } derived_variables: fopen: equation: { rhs: "1" } label: "select:gates[*]/fcond reduce:multiply" g: equation: { rhs: "conductance * fopen" } unit: nS components: n: name: n label: customGateHHrates iri: "extends:baseGate" parameters: instances: { value: 4 } state_variables: q: equation: { rhs: "(inf - q) / tau" } initial_value: 0.32 derived_variables: alpha: equation: { rhs: "0" } label: "select:forwardRate/r" unit: per_ms beta: equation: { rhs: "0" } label: "select:reverseRate/r" unit: per_ms inf: equation: { rhs: "alpha / (alpha + beta)" } tau: equation: { rhs: "1 / (alpha + beta)" } unit: ms fcond: equation: { rhs: "q ** instances" } components: forwardRate: name: forwardRate label: customHHExpLinearRate iri: "extends:baseVoltageDepRate" parameters: rate: { value: 0.1, unit: per_ms } midpoint: { value: -55, unit: mV } scale: { value: 10, unit: mV } derived_variables: x: equation: { rhs: "(v - midpoint) / scale" } r: conditional: true cases: - condition: "x != 0" equation: { rhs: "rate * x / (1 - exp(-x))" } - condition: "True" equation: { rhs: "rate" } reverseRate: name: reverseRate label: customHHExpRate iri: "extends:baseVoltageDepRate" parameters: rate: { value: 0.125, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -80, unit: mV } derived_variables: r: equation: { rhs: "rate * exp((v - midpoint) / scale)" } unit: per_ms integration: step_size: 0.01 duration: 150.0 time_scale: ms """ )print (f"Model: { exp_custom. dynamics. name} " )print (f"IRI: { exp_custom. dynamics. iri} " )print (f"Components: { list (exp_custom.dynamics.components.keys())} " )
Model: HodgkinHuxleyCustom
IRI: extends:baseCellMembPot
Components: ['k', 'na', 'passive']
This version uses iri: extends:baseCellMembPot instead of iri: neuroml:pointCellCondBased. Every equation — rate functions, gating kinetics, channel conductance — is defined explicitly in the YAML. The adapter generates custom LEMS ComponentType definitions that extend base infrastructure types, achieving exact numerical identity with the standard version.
7. Run Custom Version
= exp_custom.run("neuroml" )= result_custom.integration.dataprint (f"Custom: { da_custom. dims} , shape= { da_custom. shape} " )
Custom: ('time', 'variable'), shape=(15001, 1)
8. Compare Custom vs Reference
plot_lems_comparison(“LEMS_NML2_Ex1_HH.xml”, ref_outputs, result_custom.integration.data, title_prefix=“Ex1 Custom”)