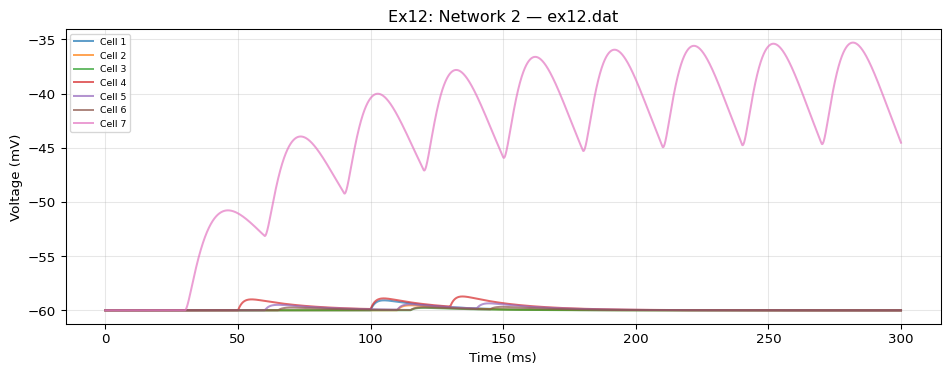
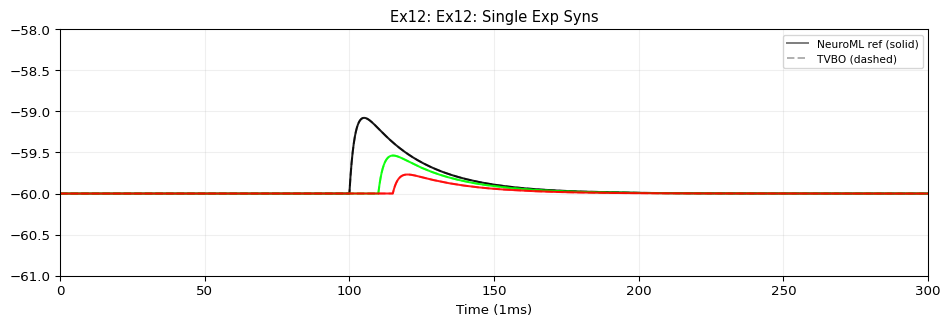
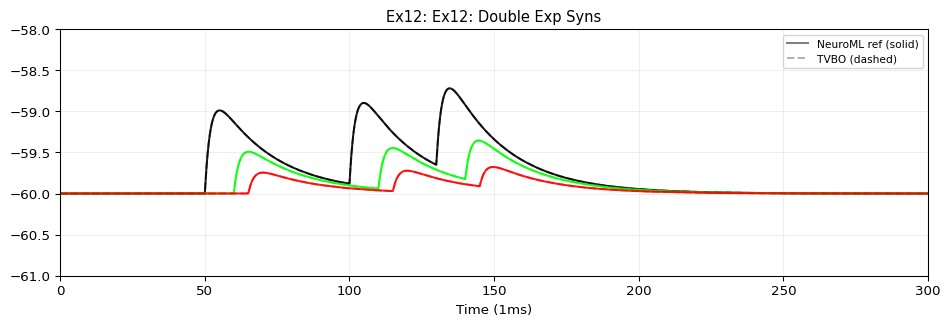
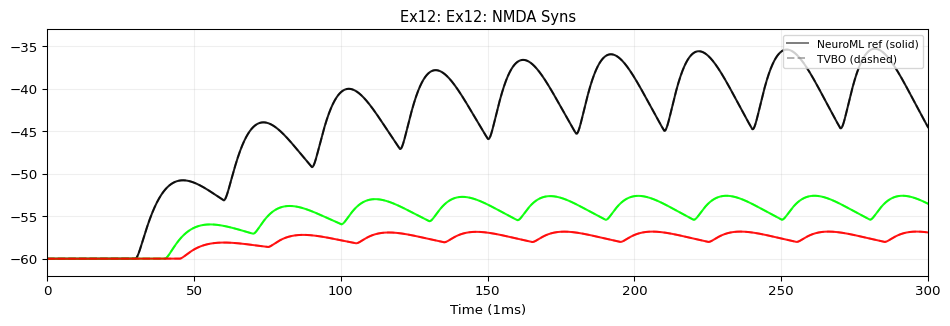
from tvbo import SimulationExperiment= SimulationExperiment.from_string(""" label: "NeuroML Ex12: Network 2" dynamics: name: iafCell iri: neuroml:iafCell network: dynamics: iaf1: name: iaf1 iri: neuroml:iafCell parameters: leakReversal: {value: -60, unit: mV} thresh: {value: -35, unit: mV} reset: {value: -65, unit: mV} C: {value: 10, unit: pF} leakConductance: {value: 0.5, unit: nS} spiker: name: spiker iri: neuroml:spikeGenerator parameters: period: {value: 30, unit: ms} spikes2: name: spikes2 iri: neuroml:spikeArray events: s: event_type: preset_time trigger_times: [50, 100, 130] spike100: name: spike100 iri: neuroml:spikeArray events: s: event_type: preset_time trigger_times: [100] # NMDA synapse with Mg²⁺ voltage-dependent block (defined with full # composition tree — blockMechanism is a child dynamics) synNmda: name: synNmda iri: neuroml:blockingPlasticSynapse parameters: gbase: {value: 5, unit: nS} tauDecay: {value: 10, unit: ms} tauRise: {value: 1, unit: ms} erev: {value: 0, unit: mV} modes: blockMechanism: name: blockMechanism iri: neuroml:voltageConcDepBlockMechanism parameters: species: {description: mg} blockConcentration: {value: 1.2, unit: mmol_per_m3} scalingConc: {value: 1.9205441817997078, unit: mmol_per_m3} scalingVolt: {value: 16.129032258064516, unit: mV} nodes: # 9 post-synaptic cells - {id: 0, dynamics: iaf1} - {id: 1, dynamics: iaf1} - {id: 2, dynamics: iaf1} - {id: 3, dynamics: iaf1} - {id: 4, dynamics: iaf1} - {id: 5, dynamics: iaf1} - {id: 6, dynamics: iaf1} - {id: 7, dynamics: iaf1} - {id: 8, dynamics: iaf1} # Spike sources - {id: 100, dynamics: spike100} - {id: 101, dynamics: spikes2} - {id: 102, dynamics: spiker} edges: # spike100 → iaf[0..2]: expOneSynapse, varying weight/delay - {source: 100, target: 0, coupling: expOneSynapse, parameters: {gbase: {value: 0.1, unit: nS}, erev: {value: 0, unit: mV}, tauDecay: {value: 2, unit: ms }} } - {source: 100, target: 1, coupling: expOneSynapse, parameters: {gbase: {value: 0.1, unit: nS}, erev: {value: 0, unit: mV}, tauDecay: {value: 2, unit: ms}, weight: {value: 0.5} , delay: {value: 10, unit: ms }} } - {source: 100, target: 2, coupling: expOneSynapse, parameters: {gbase: {value: 0.1, unit: nS}, erev: {value: 0, unit: mV}, tauDecay: {value: 2, unit: ms}, weight: {value: 0.25} , delay: {value: 15, unit: ms }} } # spikes2 → iaf[3..5]: expTwoSynapse, varying weight/delay - {source: 101, target: 3, coupling: expTwoSynapse, parameters: {gbase: {value: 0.1, unit: nS}, erev: {value: 0, unit: mV}, tauDecay: {value: 2, unit: ms}, tauRise: {value: 0.05, unit: ms }} } - {source: 101, target: 4, coupling: expTwoSynapse, parameters: {gbase: {value: 0.1, unit: nS}, erev: {value: 0, unit: mV}, tauDecay: {value: 2, unit: ms}, tauRise: {value: 0.05, unit: ms}, weight: {value: 0.5} , delay: {value: 10, unit: ms }} } - {source: 101, target: 5, coupling: expTwoSynapse, parameters: {gbase: {value: 0.1, unit: nS}, erev: {value: 0, unit: mV}, tauDecay: {value: 2, unit: ms}, tauRise: {value: 0.05, unit: ms}, weight: {value: 0.25} , delay: {value: 15, unit: ms }} } # spiker → iaf[6..8]: NMDA synapse, varying weight/delay - {source: 102, target: 6, coupling: synNmda} - {source: 102, target: 7, coupling: synNmda, parameters: {weight: {value: 0.5} , delay: {value: 10, unit: ms }} } - {source: 102, target: 8, coupling: synNmda, parameters: {weight: {value: 0.25} , delay: {value: 15, unit: ms }} } integration: method: euler step_size: 0.005 duration: 300.0 time_scale: ms """ )print (f"Model: { exp. dynamics. name if exp. dynamics else 'network' } " )