import numpy as np
import matplotlib.pyplot as plt
from tvbo import Observation, NetworkTVB Monitor Round-Trip
TVB uses monitors to define what a simulation records — raw state variables, temporal averages, BOLD fMRI via hemodynamic response, or scalp EEG/MEG via lead-field projection. TVBO stores each monitor as a framework-agnostic Observation YAML that captures the full signal processing pipeline, then converts to any backend on demand.
1. Observation Model Overview
Every observation model lives in tvbo/database/observation_models/ as a self-describing YAML file. Each defines:
- name / label — identifier and human-readable name
- class_reference — maps to a specific TVB monitor class
- period — sampling interval in ms
- pipeline — ordered signal processing steps (gain projection, HRF convolution, etc.)
- data_source — optional link to a sensor network (for EEG/MEG/iEEG)
- parameters — physical constants (conductivity, permeability, TR, …)
print("Available observation models:")
for name in Observation.list_db():
obs = Observation.from_db(name)
label = obs.label or obs.name
pipe = len(obs.pipeline or [])
ds = "sensor network" if obs.data_source else "—"
print(f" {str(obs.name):40s} pipeline={pipe} data_source={ds}")Available observation models:
AfferentCoupling pipeline=0 data_source=—
AfferentCouplingTemporalAverage pipeline=1 data_source=—
BOLD_DoubleExponential pipeline=4 data_source=—
BOLD_Gamma pipeline=4 data_source=—
BOLD_MixtureOfGammas pipeline=4 data_source=—
BOLD_RegionROI pipeline=6 data_source=—
BOLD_TVB pipeline=5 data_source=—
EEG pipeline=3 data_source=sensor network
FunctionalConnectivity pipeline=4 data_source=—
GlobalAverage pipeline=1 data_source=—
MEG pipeline=3 data_source=sensor network
Raw pipeline=0 data_source=—
SpatialAverage pipeline=1 data_source=—
SubSample pipeline=1 data_source=—
TemporalAverage pipeline=1 data_source=—
iEEG pipeline=3 data_source=sensor networkThe models fall into four categories:
| Category | Models | Key Feature |
|---|---|---|
| Basic | Raw, SubSample, TemporalAverage, GlobalAverage, SpatialAverage | Temporal/spatial reduction only |
| Projection | EEG, MEG, iEEG | Lead-field gain matrix + sensor network |
| Hemodynamic | BOLD (4 HRF variants), BoldRegionROI | HRF convolution + Volterra transform |
| Coupling | AfferentCoupling, AfferentCouplingTemporalAverage | Records coupling input, not state |
2. Basic Monitors
Basic monitors apply simple temporal or spatial transformations to the raw simulation state. They require no external data.
obs_raw = Observation.from_db("raw")
print(f"{obs_raw.name}: {obs_raw.description}")
print(f" Pipeline steps: {len(obs_raw.pipeline or [])}")
print(f" TVB class: {obs_raw.class_reference.name}")Raw: Records all state variables at every integration step without any transformation. Identity passthrough of the full simulation state.
Pipeline steps: 0
TVB class: Rawobs_ta = Observation.from_db("temporal_average")
print(f"{obs_ta.name}: {obs_ta.label}")
print(f" Period: {obs_ta.period} ms")
print(f" Pipeline: {[str(s.name) for s in obs_ta.pipeline]}")TemporalAverage: Temporal Average
Period: 0.9765625 ms
Pipeline: ['temporal_average']Convert to TVB
Each observation converts to its corresponding TVB monitor with a single function call:
from tvbo.adapters.tvb import to_tvb_monitor
mon_raw = to_tvb_monitor(obs_raw)
mon_ta = to_tvb_monitor(obs_ta)
print(f"Raw → {type(mon_raw).__name__}")
print(f"TA → {type(mon_ta).__name__}, period={mon_ta.period}")Raw → Raw
TA → TemporalAverage, period=0.97656253. Projection Monitors — EEG, MEG, iEEG
Projection monitors transform source-level neural activity into sensor signals via a lead-field (gain) matrix. Each model links to a sensor network stored as a YAML + HDF5 pair in the tvbo database.
The forward model pipeline:
- Compute gain — load precomputed lead-field or use analytic formula (Sarvas 1987)
- Lead-field projection — multiply gain matrix with source activity
- Post-processing — re-referencing (EEG) or noise addition (MEG, iEEG)
3.1 EEG
obs_eeg = Observation.from_db("eeg")
print(f"{obs_eeg.name}: {obs_eeg.label}")
print(f" Imaging modality: {obs_eeg.imaging_modality}")
print(f" Sensor network: {obs_eeg.data_source.path}")
print(f" Pipeline: {[str(s.name) for s in obs_eeg.pipeline]}")
# Parameters
for pname, p in obs_eeg.parameters.items():
unit = getattr(p, 'unit', '') or ''
val = getattr(p, 'value', '—') or '—'
print(f" {pname}: {val} {unit}")EEG: Scalp EEG
Imaging modality: EEG
Sensor network: acq-EEGBrainstorm65_sensors.yaml
Pipeline: ['compute_gain', 'lead_field_projection', 'rereference']
conductivity: 1.0 S_per_m
reference_electrode: — # Load the sensor network from database
from tvbo import database_path
sensor_net = Network.from_file(
database_path / "networks" / obs_eeg.data_source.path
)
gain = sensor_net.matrix("gain", format="dense")
print(f"Sensor network: {sensor_net.number_of_nodes} electrodes")
print(f"Gain matrix: {gain.shape} (sensors × vertices)")
print(f"Gain range: [{np.nanmin(gain):.2f}, {np.nanmax(gain):.2f}]")Sensor network: 65 electrodes
Gain matrix: (65, 16384) (sensors × vertices)
Gain range: [-120.12, 253.93]3.2 MEG
obs_meg = Observation.from_db("meg")
print(f"{obs_meg.name}: {obs_meg.label}")
print(f" Sensor network: {obs_meg.data_source.path}")
print(f" Pipeline: {[str(s.name) for s in obs_meg.pipeline]}")
meg_net = Network.from_file(
database_path / "networks" / obs_meg.data_source.path
)
meg_gain = meg_net.matrix("gain", format="dense")
print(f" {meg_net.number_of_nodes} sensors, gain {meg_gain.shape}")MEG: MEG
Sensor network: acq-MEGBrainstorm276_sensors.yaml
Pipeline: ['compute_gain', 'lead_field_projection', 'add_noise']
276 sensors, gain (276, 16384)3.3 iEEG (SEEG)
obs_ieeg = Observation.from_db("ieeg")
print(f"{obs_ieeg.name}: {obs_ieeg.label}")
print(f" Sensor network: {obs_ieeg.data_source.path}")
print(f" Pipeline: {[str(s.name) for s in obs_ieeg.pipeline]}")iEEG: Intracranial EEG (SEEG)
Sensor network: acq-SEEG588_sensors.yaml
Pipeline: ['compute_gain', 'lead_field_projection', 'add_noise']Convert Projection Monitors to TVB
The adapter loads sensors, orientations (MEG), and gain matrices automatically from the linked sensor network:
mon_eeg = to_tvb_monitor(obs_eeg)
mon_meg = to_tvb_monitor(obs_meg)
mon_ieeg = to_tvb_monitor(obs_ieeg)
print(f"EEG → {type(mon_eeg).__name__}")
print(f" Sensors: {type(mon_eeg.sensors).__name__} {mon_eeg.sensors.locations.shape}")
print(f" Projection: {type(mon_eeg.projection).__name__} {mon_eeg.projection.projection_data.shape}")
print(f"MEG → {type(mon_meg).__name__}")
print(f" Sensors: {type(mon_meg.sensors).__name__} {mon_meg.sensors.locations.shape}")
print(f" Projection: {type(mon_meg.projection).__name__} {mon_meg.projection.projection_data.shape}")
print(f"iEEG → {type(mon_ieeg).__name__}")
print(f" Sensors: {type(mon_ieeg.sensors).__name__} {mon_ieeg.sensors.locations.shape}")EEG → EEG
Sensors: SensorsEEG (65, 3)
Projection: ProjectionSurfaceEEG (65, 16384)
MEG → MEG
Sensors: SensorsMEG (276, 3)
Projection: ProjectionSurfaceMEG (276, 16384)
iEEG → iEEG
Sensors: SensorsInternal (588, 3)4. Visualize TVB Projection Networks on TVB Surface
We now build sensor-to-region projection graphs directly from the TVB-default EEG/MEG/iEEG monitor data and render them on the TVB default cortical surface mesh.
import nibabel as nib
from bsplot.graph import create_network, plot_network_on_surface
from tvbo import database_path
from tvb.datatypes.connectivity import Connectivity
from tvb.datatypes.surfaces import CorticalSurface
from tvb.datatypes.region_mapping import RegionMapping
# TVB default region/surface data
conn = Connectivity.from_file()
conn.configure()
surf = CorticalSurface.from_file()
surf.configure()
rmap = RegionMapping.from_file()
rmap.connectivity = conn
rmap.surface = surf
rmap.configure()
region_labels = list(conn.region_labels)
region_centers = conn.centres
n_regions = len(region_labels)
# Build a Gifti surface from TVB default cortical mesh (no template/MNI surface)
tvb_surface_gifti = nib.gifti.GiftiImage()
tvb_surface_gifti.add_gifti_data_array(
nib.gifti.GiftiDataArray(
np.asarray(surf.vertices, dtype=np.float32),
intent="NIFTI_INTENT_POINTSET",
)
)
tvb_surface_gifti.add_gifti_data_array(
nib.gifti.GiftiDataArray(
np.asarray(surf.triangles, dtype=np.int32),
intent="NIFTI_INTENT_TRIANGLE",
)
)
print(f"TVB surface: {surf.vertices.shape[0]} vertices, {surf.triangles.shape[0]} triangles")
print(f"TVB regions: {n_regions}")TVB surface: 16384 vertices, 32760 triangles
TVB regions: 76def _to_region_gain(gain_matrix, mapping, n_regions):
gain_abs = np.abs(np.nan_to_num(gain_matrix, nan=0.0))
# Already sensors × regions
if gain_abs.shape[1] == n_regions:
return gain_abs
# Sensors × vertices -> aggregate using region mapping
if gain_abs.shape[1] == len(mapping):
out = np.zeros((gain_abs.shape[0], n_regions), dtype=float)
for r in range(n_regions):
mask = mapping == r
if mask.any():
out[:, r] = gain_abs[:, mask].mean(axis=1)
return out
raise ValueError(
f"Unsupported gain shape {gain_abs.shape}; expected (*, {n_regions}) "
f"or (*, {len(mapping)})"
)
def _build_projection_graph(obs_name, layer_name, threshold_percentile=97):
obs = Observation.from_db(obs_name)
net = Network.from_file(database_path / "networks" / obs.data_source.path)
sensor_labels = [str(n.label) for n in net.nodes]
sensor_positions = np.asarray(
[[n.position.x, n.position.y, n.position.z] for n in net.nodes],
dtype=float,
)
n_sensors = len(sensor_labels)
gain = net.matrix("gain", format="dense")
if gain is None:
return None, None # No stored gain matrix (e.g. SEEG)
region_gain = _to_region_gain(gain, rmap.array_data, n_regions)
centers = {}
for i, lbl in enumerate(region_labels):
centers[f"R:{lbl}"] = tuple(region_centers[i])
for i, lbl in enumerate(sensor_labels):
centers[f"S:{lbl}"] = tuple(sensor_positions[i])
combined = np.zeros((n_regions + n_sensors, n_regions + n_sensors), dtype=float)
for i in range(n_sensors):
for j in range(n_regions):
combined[n_regions + i, j] = region_gain[i, j]
labels = [f"R:{x}" for x in region_labels] + [f"S:{x}" for x in sensor_labels]
graph = create_network(
centers,
{layer_name: combined},
labels=labels,
threshold_percentile=threshold_percentile,
directed=True,
edge_data_key="gain",
)
for node in graph.nodes():
graph.nodes[node]["color"] = "gold" if node.startswith("R:") else "crimson"
return graph, region_gaincandidates = {
"EEG": ("eeg", "eeg_projection"),
"MEG": ("meg", "meg_projection"),
"iEEG/SEEG": ("ieeg", "ieeg_projection"),
}
graphs = {}
for name, (obs_name, layer) in candidates.items():
result = _build_projection_graph(obs_name, layer)
if result[0] is None:
print(f"{name:10s} — no stored gain matrix, skipped")
else:
g, rg = result
graphs[name] = (g, rg)
print(
f"{name:10s} nodes={g.number_of_nodes():3d} edges={g.number_of_edges():5d} "
f"gain=[{rg.min():.2e}, {rg.max():.2e}]"
)EEG nodes=141 edges= 144 gain=[0.00e+00, 5.49e+01]
MEG nodes=352 edges= 566 gain=[0.00e+00, 9.42e-06]
iEEG/SEEG — no stored gain matrix, skippedn_plots = len(graphs)
fig, axes = plt.subplots(1, n_plots, figsize=(5 * n_plots, 5))
if n_plots == 1:
axes = [axes]
for ax, (title, (graph, _)) in zip(axes, graphs.items()):
plot_network_on_surface(
graph,
ax=ax,
template=tvb_surface_gifti,
hemi="lh",
view="top",
node_radius=1.4,
node_color="auto",
edge_radius=0.003,
edge_color="auto",
edge_data_key="gain",
edge_cmap="viridis",
edge_scale={"gain": 10, "mode": "quantile"},
surface_alpha=0.12,
)
ax.set_title(f"{title} ({graph.number_of_edges()} edges)")
plt.tight_layout()
plt.show()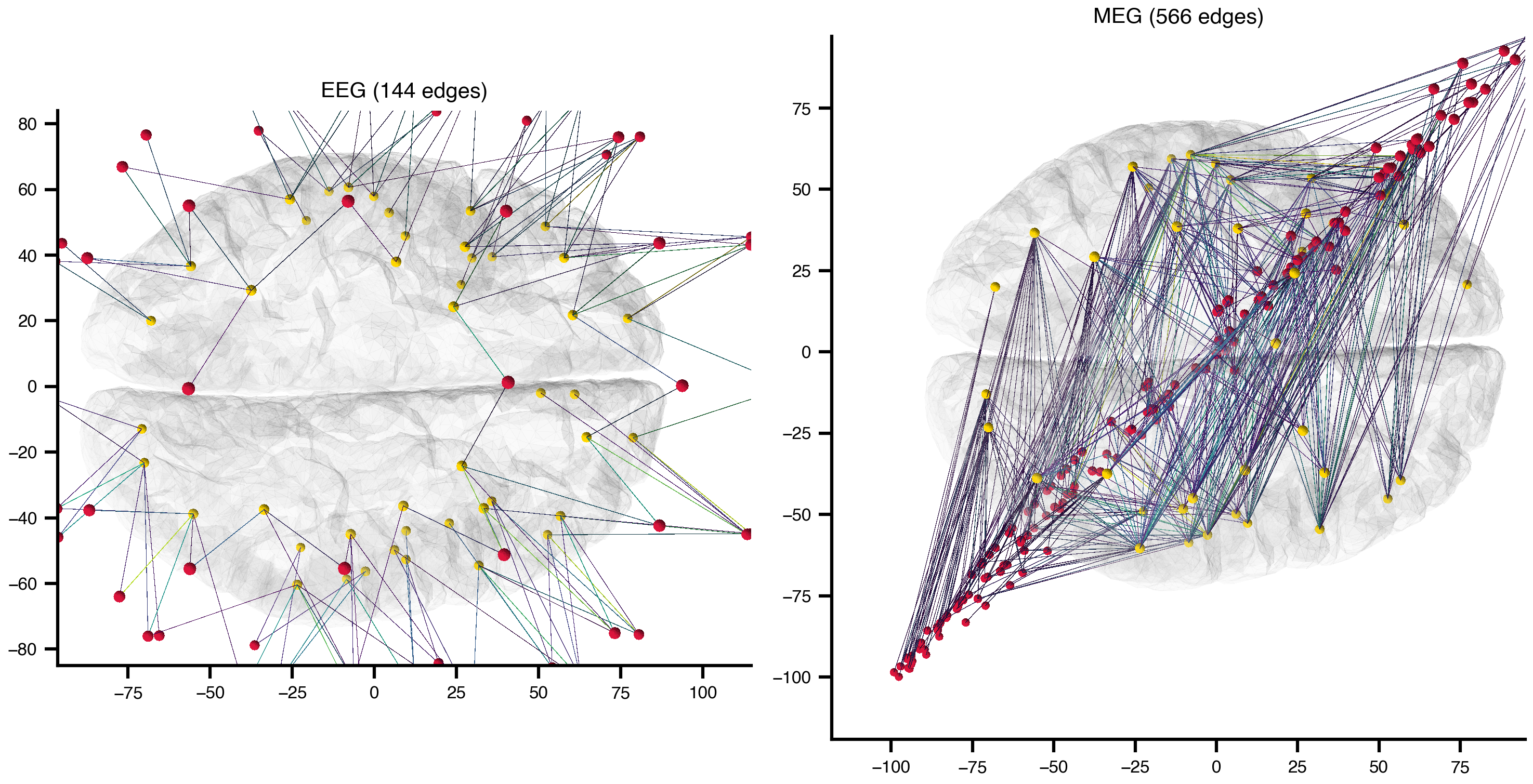
5. Hemodynamic Monitors (BOLD)
BOLD monitors simulate the fMRI signal by convolving neural activity with a hemodynamic response function (HRF), followed by temporal subsampling at the repetition time (TR).
The pipeline:
- Temporal averaging — interim downsampling before convolution
- HRF generation — compute the kernel (Volterra, Gamma, DoubleExponential, MixtureOfGammas)
- Convolution —
scipy.signal.fftconvolvewith HRF kernel - Subsampling — downsample to TR
- Volterra transform — nonlinear signal transformation (some variants)
obs_bold = Observation.from_db("bold_tvb")
print(f"{obs_bold.name}: {obs_bold.label}")
print(f" HRF kernel: {obs_bold.class_reference.constructor_args[0].value}")
print(f" Pipeline: {[str(s.name) for s in obs_bold.pipeline]}")
# Show HRF equation
hrf_step = [s for s in obs_bold.pipeline if str(s.name) == 'hemodynamic_response'][0]
print(f" HRF equation: {hrf_step.equation.rhs}")
for pname, p in hrf_step.equation.parameters.items():
print(f" {pname} = {p.value} {getattr(p, 'unit', '')}")BOLD_TVB: BOLD (First Order Volterra)
HRF kernel: FirstOrderVolterra
Pipeline: ['temporal_average_interim', 'hemodynamic_response', 'convolve', 'subsample_to_period', 'volterra_transform']
HRF equation: 1/3 * exp(-0.5*(t / tau_s)) * sin(sqrt(1/tau_f - 1/(4*tau_s**2)) * t) / sqrt(1/tau_f - 1/(4*tau_s**2))
tau_f = 0.4 s
tau_s = 0.8 smon_bold = to_tvb_monitor(obs_bold)
print(f"BOLD → {type(mon_bold).__name__}")
print(f" HRF: {type(mon_bold.hrf_kernel).__name__}")
print(f" Period: {mon_bold.period} ms")
# Available BOLD variants
bold_variants = [n for n in Observation.list_db() if 'bold' in n.lower()]
for name in bold_variants:
obs = Observation.from_db(name)
cr = obs.class_reference
hrf = next((a.value for a in (cr.constructor_args or []) if str(a.name) == 'hrf_kernel'), '—')
print(f" {str(obs.name):30s} HRF={hrf}")BOLD → Bold
HRF: FirstOrderVolterra
Period: 2000.0 ms
BOLD_DoubleExponential HRF=DoubleExponential
BOLD_Gamma HRF=Gamma
BOLD_MixtureOfGammas HRF=MixtureOfGammas
BOLD_RegionROI HRF=FirstOrderVolterra
BOLD_TVB HRF=FirstOrderVolterra6. Coupling Monitors
Coupling monitors record the afferent (incoming) coupling term at each node — the network input that drives the dynamics, rather than the model state itself. Useful for analysing effective connectivity.
obs_ac = Observation.from_db("afferent_coupling")
print(f"{obs_ac.name}: {obs_ac.label}")
print(f" Source: {obs_ac.source}")
print(f" Pipeline steps: {len(obs_ac.pipeline or [])}")
mon_ac = to_tvb_monitor(obs_ac)
print(f" → {type(mon_ac).__name__}")AfferentCoupling: Afferent Coupling Recording
Source: coupling
Pipeline steps: 0
→ AfferentCoupling7. Simulation Round-Trip: Numerical Verification
To verify that tvbo-generated TVB models are faithful to the original, we compare simulations run with the TVB-native Generic2dOscillator against the tvbo-generated equivalent. Both use identical connectivity, coupling, integrator, and initial conditions. Results are wrapped using ExperimentResult.from_tvb() for consistent access.
from tvbo import Dynamics, ExperimentResult
from tvb.simulator.lab import simulator, models, coupling, integrators, monitors
# Shared setup
sim_length = 100.0 # ms
seed = 42
# --- TVB Native ---
m1 = models.Generic2dOscillator()
m1.variables_of_interest = ('V', 'W')
sim1 = simulator.Simulator(
model=m1,
coupling=coupling.Linear(a=np.array([0.0126])),
integrator=integrators.HeunDeterministic(dt=0.1),
connectivity=conn,
monitors=[monitors.Raw()],
simulation_length=sim_length,
)
np.random.seed(seed)
sim1.configure()
result_native = ExperimentResult.from_tvb(sim1)
# --- tvbo Generated ---
m = Dynamics.from_db('Generic2dOscillator')
code = m.render_code('tvb')
ns = {}
exec(code, ns)
m2 = ns['Generic2dOscillator']()
sim2 = simulator.Simulator(
model=m2,
coupling=coupling.Linear(a=np.array([0.0126])),
integrator=integrators.HeunDeterministic(dt=0.1),
connectivity=conn,
monitors=[monitors.Raw()],
simulation_length=sim_length,
)
np.random.seed(seed)
sim2.configure()
result_tvbo = ExperimentResult.from_tvb(sim2)
print(f"TVB native cvar: {sim1.model.cvar}")
print(f"tvbo-generated cvar: {sim2.model.cvar}")
print(f"Output shape: {result_native.integration.data.shape}")
diff = np.abs(result_native.integration.data - result_tvbo.integration.data).max()
print(f"Max absolute diff: {diff:.2e}")
print(f"Numerically equal: {np.allclose(result_native.integration.data, result_tvbo.integration.data, atol=1e-10)}")TVB native cvar: [0]
tvbo-generated cvar: [0]
Output shape: (1000, 2, 76, 1)
Max absolute diff: 3.55e-15
Numerically equal: Truets_native = result_native.integration
ts_tvbo = result_tvbo.integration
node = 0
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(10, 4), sharex=True, height_ratios=[3, 1])
ts_native.get_subspace_by_index([node]).plot(ax=ax1, label="TVB native", linewidth=1.5)
ts_tvbo.get_subspace_by_index([node]).plot(ax=ax1, label="tvbo-generated", linestyle="--", linewidth=1.5)
ax1.legend()
ax1.set_title(f"Region {node}: {conn.region_labels[node]}")
diff_trace = np.abs(ts_native.data.isel(variable=0, node=node, mode=0) - ts_tvbo.data.isel(variable=0, node=node, mode=0)) + 1e-20
ax2.semilogy(ts_native.time, diff_trace, color="k", lw=0.8)
ax2.set_ylabel("|diff|")
ax2.set_xlabel("Time (ms)")
plt.tight_layout()
plt.show()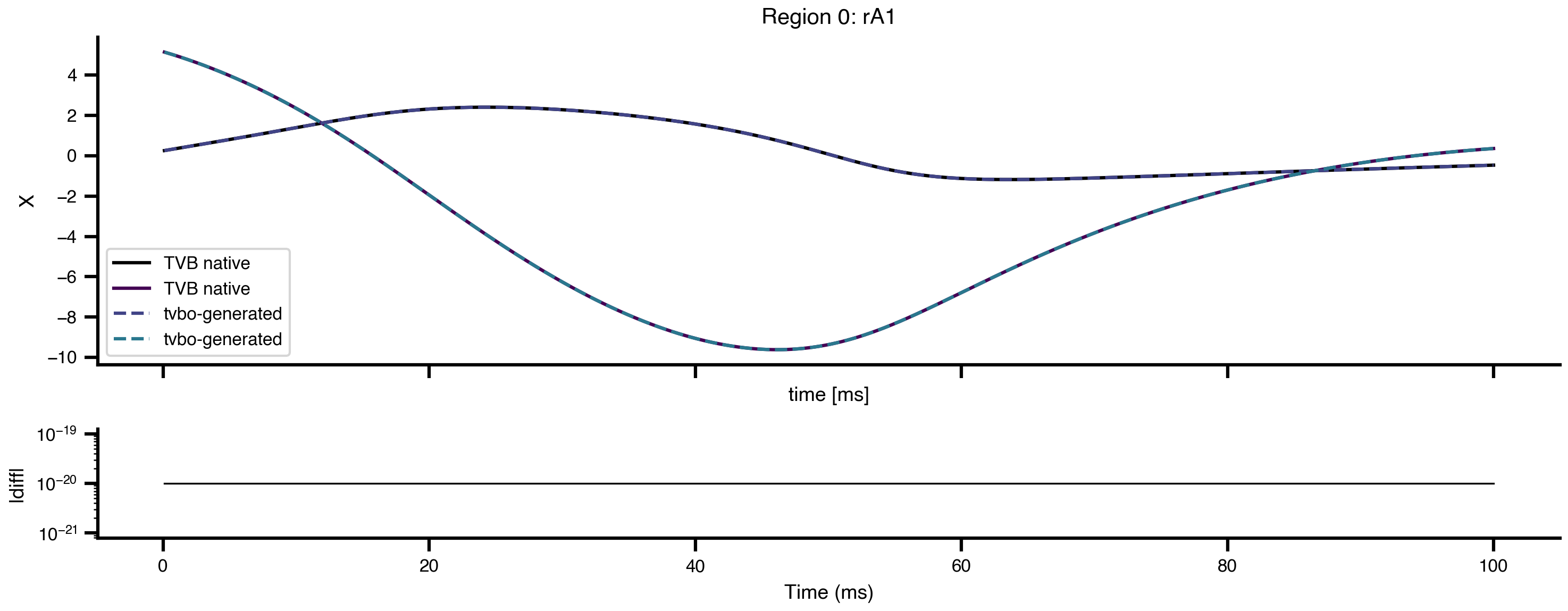
Code Generation API
Each observation model can also render its TVB monitor code directly:
obs_eeg_rt = Observation.from_db("eeg")
print(obs_eeg_rt.render_code("tvb"))import abc
import numpy
from tvb.simulator.monitors import Monitor
from tvb.simulator.monitors import Projection
from tvb.basic.neotraits.api import Float, NArray, Attr
from tvb.simulator.backend.ref import ReferenceBackend
from tvb.datatypes.sensors import SensorsEEG
from tvb.datatypes.projections import ProjectionSurfaceEEG
from tvb.simulator import noise
from tvb.datatypes.region_mapping import RegionMapping
from tvb.simulator.common import numpy_add_at
class EEG(Projection):
"""Lead-field projection monitor (EEG)."""
period = Float(default=0.9765625)
sigma = Float(default=1.0)
sensors = Attr(SensorsEEG, required=True)
projection = Attr(ProjectionSurfaceEEG, default=None)
reference = Attr(str, required=False, default=None)
def analytic(self, loc, ori):
"""Sarvas 1987 single-sphere approximation (Eq. 12)."""
r_0, Q = loc, ori
centre = numpy.mean(r_0, axis=0)[numpy.newaxis, :]
radius = 1.05125 * max(numpy.sqrt(numpy.sum((r_0 - centre) ** 2, axis=1)))
sen_loc = self.sensors.locations.copy()
sen_dis = numpy.sqrt(numpy.sum(sen_loc**2, axis=1))
sen_loc = sen_loc / sen_dis[:, numpy.newaxis] * radius + centre
V_r = numpy.zeros((sen_loc.shape[0], r_0.shape[0]))
for k in range(sen_loc.shape[0]):
a = sen_loc[k, :] - r_0
na = numpy.sqrt(numpy.sum(a**2, axis=1))[:, numpy.newaxis]
V_r[k, :] = numpy.sum(Q * (a / na**3), axis=1) / (
4.0 * numpy.pi * self.sigma
)
return V_r
def config_for_sim(self, simulator):
super(EEG, self).config_for_sim(simulator)
n_sensors = self.sensors.number_of_sensors
self._ref_vec = numpy.zeros((n_sensors,))
if self.reference:
if self.reference.lower() != "average":
idx = self.sensors.labels.tolist().index(self.reference)
self._ref_vec[idx] = 1.0
else:
self._ref_vec[:] = 1.0 / n_sensors
self._ref_vec_mask = numpy.isfinite(self.gain).all(axis=1)
self._ref_vec = self._ref_vec[self._ref_vec_mask]
def sample(self, step, state):
maybe = super(EEG, self).sample(step, state)
if maybe is not None:
time, sample = maybe
sample -= self._ref_vec.dot(sample[:, self._ref_vec_mask])[:, numpy.newaxis]
return time, sample.reshape((self.voi.size, -1, 1))
monitors = [
EEG(sensors=SensorsEEG.from_file(), projection=ProjectionSurfaceEEG.from_file()),
]
# Execute returns a configured TVB monitor object
mon = obs_eeg_rt.execute("tvb")
print(f"Returned: {type(mon).__name__}")
if hasattr(mon, 'sensors') and mon.sensors is not None:
print(f" Sensors: {mon.sensors.locations.shape}")Returned: EEG
Sensors: (65, 3)8. Monitor Round-Trip: tvbo → TVB → verify
The adapter can convert every observation type to its corresponding TVB monitor and back:
from tvbo.adapters.tvb import to_tvb_monitor
for name in ["raw", "eeg", "bold_tvb", "temporal_average"]:
obs = Observation.from_db(name)
mon = to_tvb_monitor(obs)
print(f"{str(obs.name):20s} → {type(mon).__name__:30s}", end="")
if hasattr(mon, 'sensors') and mon.sensors is not None:
print(f" sensors={mon.sensors.locations.shape}", end="")
if hasattr(mon, 'projection') and mon.projection is not None:
print(f" gain={mon.projection.projection_data.shape}", end="")
if hasattr(mon, 'hrf_kernel') and mon.hrf_kernel is not None:
print(f" hrf={type(mon.hrf_kernel).__name__}", end="")
print()Raw → Raw
EEG → EEG sensors=(65, 3) gain=(65, 16384)
BOLD_TVB → Bold hrf=FirstOrderVolterra
TemporalAverage → TemporalAverage Summary
| Step | Method | What It Does |
|---|---|---|
| List | Observation.list_db() |
Show all available observation models |
| Load | Observation.from_db("eeg") |
Load observation from database by name |
| File | Observation.from_file("path.yaml") |
Load from arbitrary YAML file |
| Export | to_tvb_monitor(obs) |
Convert to configured TVB Monitor |
| Sensors | Network.from_file(database_path / "networks" / obs.data_source.path) |
Load linked sensor network |
| Gain | sensor_net.matrix("gain", format="dense") |
Access lead-field matrix |
The observation model YAML captures the complete signal processing chain — from raw neural activity to the measured signal — in a framework-agnostic format that any backend can consume.
