from tvbo import SimulationExperiment
exp = SimulationExperiment.from_string("""
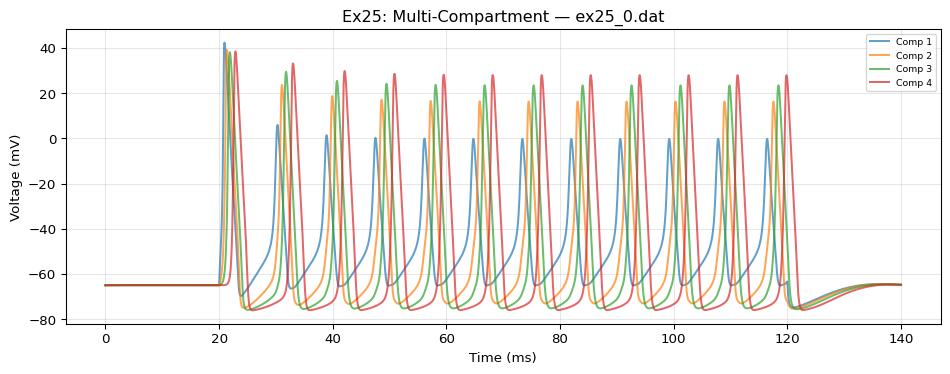
label: "NeuroML Ex25: MultiComp"
network:
dynamics:
# ── Ion Channels (imported from NML2_SingleCompHHCell.nml) ──
passiveChan:
name: passiveChan
iri: neuroml:ionChannelHH
parameters:
conductance: {value: 10, unit: pS}
condDensity: {value: 3.0, unit: S_per_m2}
erev: {value: -54.3, unit: mV}
ion: {label: non_specific}
naChan:
name: naChan
iri: neuroml:ionChannelHH
parameters:
conductance: {value: 10, unit: pS}
species: {label: na}
condDensity: {value: 120.0, unit: mS_per_cm2}
erev: {value: 50.0, unit: mV}
ion: {label: na}
modes:
m:
name: m
iri: neuroml:gateHHrates
parameters:
instances: {value: 3}
modes:
forwardRate:
name: forwardRate
iri: neuroml:HHExpLinearRate
parameters:
rate: {value: 1, unit: per_ms}
midpoint: {value: -40, unit: mV}
scale: {value: 10, unit: mV}
reverseRate:
name: reverseRate
iri: neuroml:HHExpRate
parameters:
rate: {value: 4, unit: per_ms}
midpoint: {value: -65, unit: mV}
scale: {value: -18, unit: mV}
h:
name: h
iri: neuroml:gateHHrates
parameters:
instances: {value: 1}
modes:
forwardRate:
name: forwardRate
iri: neuroml:HHExpRate
parameters:
rate: {value: 0.07, unit: per_ms}
midpoint: {value: -65, unit: mV}
scale: {value: -20, unit: mV}
reverseRate:
name: reverseRate
iri: neuroml:HHSigmoidRate
parameters:
rate: {value: 1, unit: per_ms}
midpoint: {value: -35, unit: mV}
scale: {value: 10, unit: mV}
kChan:
name: kChan
iri: neuroml:ionChannelHH
parameters:
conductance: {value: 10, unit: pS}
species: {label: k}
condDensity: {value: 360, unit: S_per_m2}
erev: {value: -77, unit: mV}
ion: {label: k}
modes:
n:
name: n
iri: neuroml:gateHHrates
parameters:
instances: {value: 4}
modes:
forwardRate:
name: forwardRate
iri: neuroml:HHExpLinearRate
parameters:
rate: {value: 0.1, unit: per_ms}
midpoint: {value: -55, unit: mV}
scale: {value: 10, unit: mV}
reverseRate:
name: reverseRate
iri: neuroml:HHExpRate
parameters:
rate: {value: 0.125, unit: per_ms}
midpoint: {value: -65, unit: mV}
scale: {value: -80, unit: mV}
# ── Multi-Compartment Cell ──
MultiCompCell:
name: MultiCompCell
iri: neuroml:cell
parameters:
specificCapacitance: {value: 1.0, unit: uF_per_cm2}
initMembPotential: {value: -65, unit: mV}
spikeThresh: {value: -20, unit: mV}
resistivity: {value: 100, unit: kohm_cm}
modes:
# Channels (shared across all segments via channelDensity)
passiveChan:
name: passiveChan
iri: neuroml:ionChannelHH
parameters:
conductance: {value: 10, unit: pS}
condDensity: {value: 3.0, unit: S_per_m2}
erev: {value: -54.3, unit: mV}
ion: {label: non_specific}
naChan:
name: naChan
iri: neuroml:ionChannelHH
parameters:
conductance: {value: 10, unit: pS}
species: {label: na}
condDensity: {value: 120.0, unit: mS_per_cm2}
erev: {value: 50.0, unit: mV}
ion: {label: na}
modes:
m:
name: m
iri: neuroml:gateHHrates
parameters:
instances: {value: 3}
modes:
forwardRate:
name: forwardRate
iri: neuroml:HHExpLinearRate
parameters:
rate: {value: 1, unit: per_ms}
midpoint: {value: -40, unit: mV}
scale: {value: 10, unit: mV}
reverseRate:
name: reverseRate
iri: neuroml:HHExpRate
parameters:
rate: {value: 4, unit: per_ms}
midpoint: {value: -65, unit: mV}
scale: {value: -18, unit: mV}
h:
name: h
iri: neuroml:gateHHrates
parameters:
instances: {value: 1}
modes:
forwardRate:
name: forwardRate
iri: neuroml:HHExpRate
parameters:
rate: {value: 0.07, unit: per_ms}
midpoint: {value: -65, unit: mV}
scale: {value: -20, unit: mV}
reverseRate:
name: reverseRate
iri: neuroml:HHSigmoidRate
parameters:
rate: {value: 1, unit: per_ms}
midpoint: {value: -35, unit: mV}
scale: {value: 10, unit: mV}
kChan:
name: kChan
iri: neuroml:ionChannelHH
parameters:
conductance: {value: 10, unit: pS}
species: {label: k}
condDensity: {value: 360, unit: S_per_m2}
erev: {value: -77, unit: mV}
ion: {label: k}
modes:
n:
name: n
iri: neuroml:gateHHrates
parameters:
instances: {value: 4}
modes:
forwardRate:
name: forwardRate
iri: neuroml:HHExpLinearRate
parameters:
rate: {value: 0.1, unit: per_ms}
midpoint: {value: -55, unit: mV}
scale: {value: 10, unit: mV}
reverseRate:
name: reverseRate
iri: neuroml:HHExpRate
parameters:
rate: {value: 0.125, unit: per_ms}
midpoint: {value: -65, unit: mV}
scale: {value: -80, unit: mV}
# ── Morphology: segments ──
Soma:
name: Soma
iri: neuroml:segment
parameters:
id: {value: 0}
proximal_x: {value: 0}
proximal_y: {value: 0}
proximal_z: {value: 0}
proximal_diameter: {value: 10}
distal_x: {value: 0}
distal_y: {value: 10}
distal_z: {value: 0}
distal_diameter: {value: 10}
Dendrite1:
name: Dendrite1
iri: neuroml:segment
parameters:
id: {value: 1}
parent: {value: 0}
proximal_x: {value: 0}
proximal_y: {value: 10}
proximal_z: {value: 0}
proximal_diameter: {value: 3}
distal_x: {value: 0}
distal_y: {value: 20}
distal_z: {value: 0}
distal_diameter: {value: 3}
Dendrite2a:
name: Dendrite2a
iri: neuroml:segment
parameters:
id: {value: 2}
parent: {value: 1}
proximal_x: {value: 0}
proximal_y: {value: 20}
proximal_z: {value: 0}
proximal_diameter: {value: 3}
distal_x: {value: 0}
distal_y: {value: 30}
distal_z: {value: 0}
distal_diameter: {value: 2.5}
Dendrite2b:
name: Dendrite2b
iri: neuroml:segment
parameters:
id: {value: 3}
parent: {value: 2}
distal_x: {value: 0}
distal_y: {value: 50}
distal_z: {value: 0}
distal_diameter: {value: 1.5}
# ── Morphology: segment groups ──
soma:
name: soma
iri: neuroml:segmentGroup
parameters:
neuroLexId: {label: sao864921383}
members: {shape: "0"}
dendSec1:
name: dendSec1
iri: neuroml:segmentGroup
parameters:
neuroLexId: {label: sao864921383}
members: {shape: "1"}
dendSec2:
name: dendSec2
iri: neuroml:segmentGroup
parameters:
neuroLexId: {label: sao864921383}
numberInternalDivisions: {value: 9}
members: {shape: "2,3"}
soma_group:
name: soma_group
iri: neuroml:segmentGroup
parameters:
includes: {shape: soma}
dendrite_group:
name: dendrite_group
iri: neuroml:segmentGroup
parameters:
includes: {shape: "dendSec1,dendSec2"}
# ── Synapses ──
AMPA:
name: AMPA
iri: neuroml:expTwoSynapse
parameters:
tauRise: {value: 3e-5, unit: s}
tauDecay: {value: 0.5e-3, unit: s}
gbase: {value: 0.3, unit: nS}
erev: {value: 0, unit: V}
NMDA:
name: NMDA
iri: neuroml:blockingPlasticSynapse
parameters:
gbase: {value: 0.8, unit: nS}
tauRise: {value: 1e-3, unit: s}
tauDecay: {value: 13.3333e-3, unit: s}
erev: {value: 0, unit: V}
modes:
blockMechanism:
name: blockMechanism
iri: neuroml:voltageConcDepBlockMechanism
parameters:
species: {label: mg}
blockConcentration: {value: 1.2, unit: mM}
scalingConc: {value: 1.9205441817997078, unit: mM}
scalingVolt: {value: 0.016129032258064516, unit: V}
# ── Inputs ──
pulseGen2:
name: pulseGen2
iri: neuroml:pulseGenerator
parameters:
delay: {value: 20, unit: ms}
duration: {value: 100, unit: ms}
amplitude: {value: 0.2, unit: nA}
pulseGen3:
name: pulseGen3
iri: neuroml:pulseGenerator
parameters:
delay: {value: 30, unit: ms}
duration: {value: 100, unit: ms}
amplitude: {value: 0.18, unit: nA}
# ── Network topology ──
nodes:
# 3 multi-compartment cells
- id: 0
dynamics: MultiCompCell
position: {x: 0, y: 0, z: 0}
- id: 1
dynamics: MultiCompCell
position: {x: 30, y: 0, z: 0}
- id: 2
dynamics: MultiCompCell
position: {x: 60, y: 0, z: 0}
# Input sources
- id: 100
dynamics: pulseGen2
- id: 101
dynamics: pulseGen3
edges:
# ── Inputs to cells ──
- source: 100
target: 0
parameters:
segmentId: {value: 0}
fractionAlong: {value: 0.5}
- source: 101
target: 2
parameters:
segmentId: {value: 0}
fractionAlong: {value: 0.5}
# ── AMPA projections: cells 0,2 → cell 1 ──
- source: 0
target: 1
coupling: AMPA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 0}
postFractionAlong: {value: 0.5}
- source: 0
target: 1
coupling: AMPA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 3}
postFractionAlong: {value: 0.3}
- source: 2
target: 1
coupling: AMPA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 0}
postFractionAlong: {value: 0.5}
- source: 2
target: 1
coupling: AMPA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 1}
postFractionAlong: {value: 0.5}
- source: 2
target: 1
coupling: AMPA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 3}
postFractionAlong: {value: 0.25}
# ── NMDA projections: cells 0,2 → cell 1 ──
- source: 0
target: 1
coupling: NMDA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 0}
postFractionAlong: {value: 0.5}
- source: 0
target: 1
coupling: NMDA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 3}
postFractionAlong: {value: 0.5}
- source: 2
target: 1
coupling: NMDA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 0}
postFractionAlong: {value: 0.5}
- source: 2
target: 1
coupling: NMDA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 1}
postFractionAlong: {value: 0.5}
- source: 2
target: 1
coupling: NMDA
parameters:
preSegmentId: {value: 0}
preFractionAlong: {value: 0.5}
postSegmentId: {value: 3}
postFractionAlong: {value: 0.25}
integration:
method: euler
step_size: 0.005
duration: 140.0
time_scale: ms
""")
print(f"Model: {exp.dynamics.name if exp.dynamics else 'network'}")