# =============================================================================
# Network Diffusion - Getting Started with NetworkDynamics.jl
# =============================================================================
# Source: https://juliadynamics.github.io/NetworkDynamics.jl/stable/generated/getting_started_with_network_dynamics/
#
# Diffusion on an undirected network. Each node state evolves as the sum of
# incoming edge flows.
# Vertex has no intrinsic dynamics; edge is antisymmetric: e = v_src - v_dst.
# =============================================================================
id: 100
label: "Network Diffusion"
description: >
Diffusion on an undirected network. Each node state evolves as the sum
of incoming edge flows. The edge computes the antisymmetric difference
between source and destination states: e = v_src - v_dst.
Recreates the NetworkDynamics.jl Getting Started tutorial.
references:
- "https://juliadynamics.github.io/NetworkDynamics.jl/stable/generated/getting_started_with_network_dynamics/"
# -- Local Dynamics ------------------------------------------------------------
dynamics:
name: Diffusion
label: "Diffusion vertex"
description: >
Node with no internal dynamics. The rate of change of the single
state variable v equals the aggregated incoming edge flow (esum).
Corresponds to the discrete Laplacian: dx_i = sum_j A_ji (x_j - x_i).
system_type: continuous
autonomous: true
parameters: {}
coupling_terms:
c_flow:
description: "Aggregated edge flow into this vertex (sum of incident edge outputs)"
state_variables:
v:
label: "Diffusion state"
description: "Abstract quantity (e.g. temperature, concentration) diffusing on the network"
equation:
rhs: "c_flow"
initial_value: 0.0
distribution:
name: Gaussian
seed: 1
domain: { lo: -3.0, hi: 3.0 }
variable_of_interest: true
# -- Network -------------------------------------------------------------------
network:
label: "20-node Barabási-Albert network"
description: >
Undirected Barabási-Albert scale-free random graph with 20 nodes
and average degree 4. Matches the original tutorial.
number_of_nodes: 20
graph_generator:
name: barabasi_albert
type: BarabasiAlbert
parameters:
k: { name: k, value: 4 }
coupling:
DiffusiveCoupling:
name: DiffusiveCoupling
label: "Diffusive Coupling"
description: >
Antisymmetric edge function: output is the difference between
source and destination vertex states.
g_dst(v_src, v_dst) = v_src - v_dst;
g_src(v_src, v_dst) = v_dst - v_src (by AntiSymmetric wrapper).
delayed: false
pre_expression:
rhs: "x_j - x_i"
description: "Discrete gradient: source state minus destination state"
# -- Integration ---------------------------------------------------------------
integration:
description: "Explicit Runge-Kutta (Tsit5) with dt=0.01 over t in [0, 2]"
method: Tsit5
step_size: 0.01
duration: 2.0
time_scale: "a.u."
Network Diffusion
Getting started with NetworkDynamics.jl via tvbo
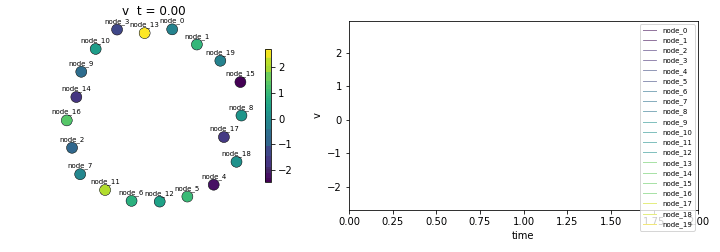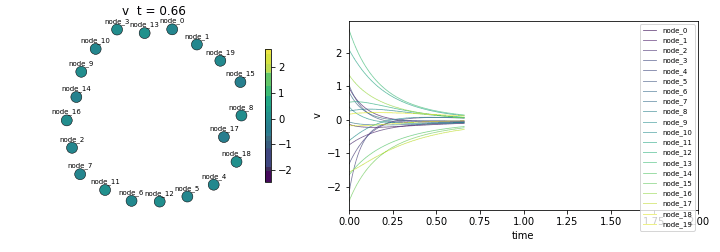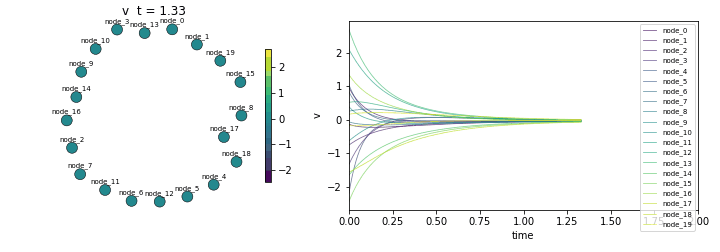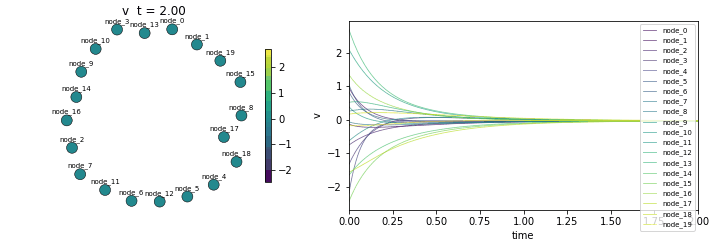
Replicates the NetworkDynamics.jl Getting Started tutorial — diffusion on an undirected network with 20 nodes. A single scalar state per node, antisymmetric diffusive coupling, and convergence to equilibrium.
YAML Specification
Model Report
Network Diffusion
Diffusion on an undirected network. Each node state evolves as the sum of incoming edge flows. The edge computes the antisymmetric difference between source and destination states: e = v_src - v_dst. Recreates the NetworkDynamics.jl Getting Started tutorial.
1. Brain Network: 20-node Barabási-Albert network
Undirected Barabási-Albert scale-free random graph with 20 nodes and average degree 4. Matches the original tutorial.
- Regions: 20
Coupling: Diffusive Coupling
Antisymmetric edge function: output is the difference between source and destination vertex states. g_dst(v_src, v_dst) = v_src - v_dst; g_src(v_src, v_dst) = v_dst - v_src (by AntiSymmetric wrapper).
Pre-synaptic: \(c_{\text{pre}} = x_{j} - x_{i}\)
2. Local Dynamics: Diffusion vertex
Node with no internal dynamics. The model comprises 1 state variables.
2.1 State Equations
\[\dot{v} = c_{flow}\]
2.2 Parameters
| Parameter | Value | Unit | Description |
|---|
3. Numerical Integration
- Method: Tsit5
- Time step: \(\Delta t = 0.01\) ms
- Duration: 2.0 ms
References
- https://juliadynamics.github.io/NetworkDynamics.jl/stable/generated/getting_started_with_network_dynamics/
Generated Julia Code
from tvbo import SimulationExperiment
exp = SimulationExperiment.from_file("yaml/diffusion.yaml")
Code(exp.render_code("networkdynamics"), language='julia')using Graphs
using NetworkDynamics
using OrdinaryDiffEqTsit5
function Diffusion_f!(dx, x, esum, p, t)
v = x[1]
c_flow = esum[1]
dx[1] = c_flow
nothing
end
vertex_Diffusion = VertexModel(;
f = Diffusion_f!,
g = StateMask(1:1),
sym = [:v],
name = :Diffusion,
)
function DiffusiveCoupling_edge_g!(e_dst, v_src, v_dst, p, t)
e_dst[1] = v_src[1] - v_dst[1]
nothing
end
edge_DiffusiveCoupling = EdgeModel(;
g = AntiSymmetric(DiffusiveCoupling_edge_g!),
outsym = [:coupling],
name = :DiffusiveCoupling,
)
g = barabasi_albert(20, 4)
nw = Network(g, vertex_Diffusion, edge_DiffusiveCoupling)
s = NWState(nw)
using Random
rng = MersenneTwister(1)
for node in 1:nv(g)
s.v[node, :v] = (-3.0 + 3.0) / 2 .+ (3.0 - -3.0) / 6 .* randn(rng)
end
tspan = (0.0, 2.0)
prob = ODEProblem(nw, uflat(s), tspan, pflat(s))
sol = solve(prob, Tsit5(); saveat=0.01)
adj_matrix = Float64.(adjacency_matrix(g))
using Random: MersenneTwister
function spring_layout(g; seed=42, iterations=50, k=1.0)
rng = MersenneTwister(seed)
n = nv(g)
pos = randn(rng, n, 2)
for _ in 1:iterations
disp = zeros(n, 2)
for i in 1:n, j in (i+1):n
d = pos[i, :] - pos[j, :]
dist = max(norm(d), 0.01)
rep = k^2 / dist
disp[i, :] .+= d / dist * rep
disp[j, :] .-= d / dist * rep
end
for e in edges(g)
i, j = src(e), dst(e)
d = pos[j, :] - pos[i, :]
dist = max(norm(d), 0.01)
att = dist^2 / k
disp[i, :] .+= d / dist * att
disp[j, :] .-= d / dist * att
end
for i in 1:n
dl = max(norm(disp[i, :]), 0.01)
pos[i, :] .+= disp[i, :] / dl * min(dl, 0.1)
end
end
return pos
end
using LinearAlgebra: norm
node_positions = spring_layout(g)
using Plots
plot(sol; ylabel="state", xlabel="time", title="Diffusion on 20-node network")
Run & Plot
ts = exp.run(format="networkdynamics")
ts.plot()Detected IPython. Loading juliacall extension. See https://juliapy.github.io/PythonCall.jl/stable/compat/#IPython
Results
Shape: (201, 1, 20)
Time range: 0.0 – 2.0
Final state std: 0.003375
(Should converge to ~0 as all nodes reach equilibrium)Animate
ani = ts.animate("v", format="dots")
ani.save("_output/diffusion.gif", writer="pillow", fps=20)