import numpy as np
import tempfile
from pathlib import Path
from tvbo import Network
from bsplot.surface import plot_surf
from bsplot.graph import create_networkTVB Connectome Round-Trip
TVB (The Virtual Brain) stores brain connectivity as Connectivity objects with weights, tract lengths, centres, region labels, and conduction speed. TVBO can import these losslessly, persist them as YAML + HDF5, and export back to TVB with identical arrays.
1. Import TVB Connectivity
Load the default TVB connectivity (76-region Hagmann parcellation) and convert it to a tvbo Network:
from tvb.datatypes.connectivity import Connectivity
conn = Connectivity.from_file()
conn.configure()
print(f"TVB: {conn.weights.shape[0]} regions, "
f"speed = {conn.speed[0]} mm/ms")TVB: 76 regions, speed = 3.0 mm/msnet = Network.from_tvb(conn)
print(f"tvbo: {net.number_of_nodes} nodes")
print(f"Label: {net.label}")
print(f"Conduction speed: {net.conduction_speed.value} {net.conduction_speed.unit}")
print(f"Weights range: [{net.weights_matrix.min():.4f}, {net.weights_matrix.max():.4f}]")
print(f"Lengths range: [{net.lengths_matrix.min():.1f}, {net.lengths_matrix.max():.1f}] mm")tvbo: 76 nodes
Label: Connectivity gid: 9df860e7-5404-4cf5-98d7-500e11dfbbed
Conduction speed: 3.0 mm_per_ms
Weights range: [0.0000, 3.0000]
Lengths range: [0.0, 153.5] mmAll TVB per-node metadata is preserved — cortical flags, areas, hemispheres, and orientations:
node0 = net.nodes[0]
print(f"Node 0: {node0.label}")
print(f" Position: ({node0.position.x:.1f}, {node0.position.y:.1f}, {node0.position.z:.1f})")
if node0.parameters:
params = node0.parameters
for name in ["cortical", "area", "hemisphere"]:
p = getattr(params, name, None)
if p is not None:
print(f" {name}: {p.value}")Node 0: rA1
Position: (-9.9, -47.1, -3.1)2. Visualize the Connectome
net.plot_matrix()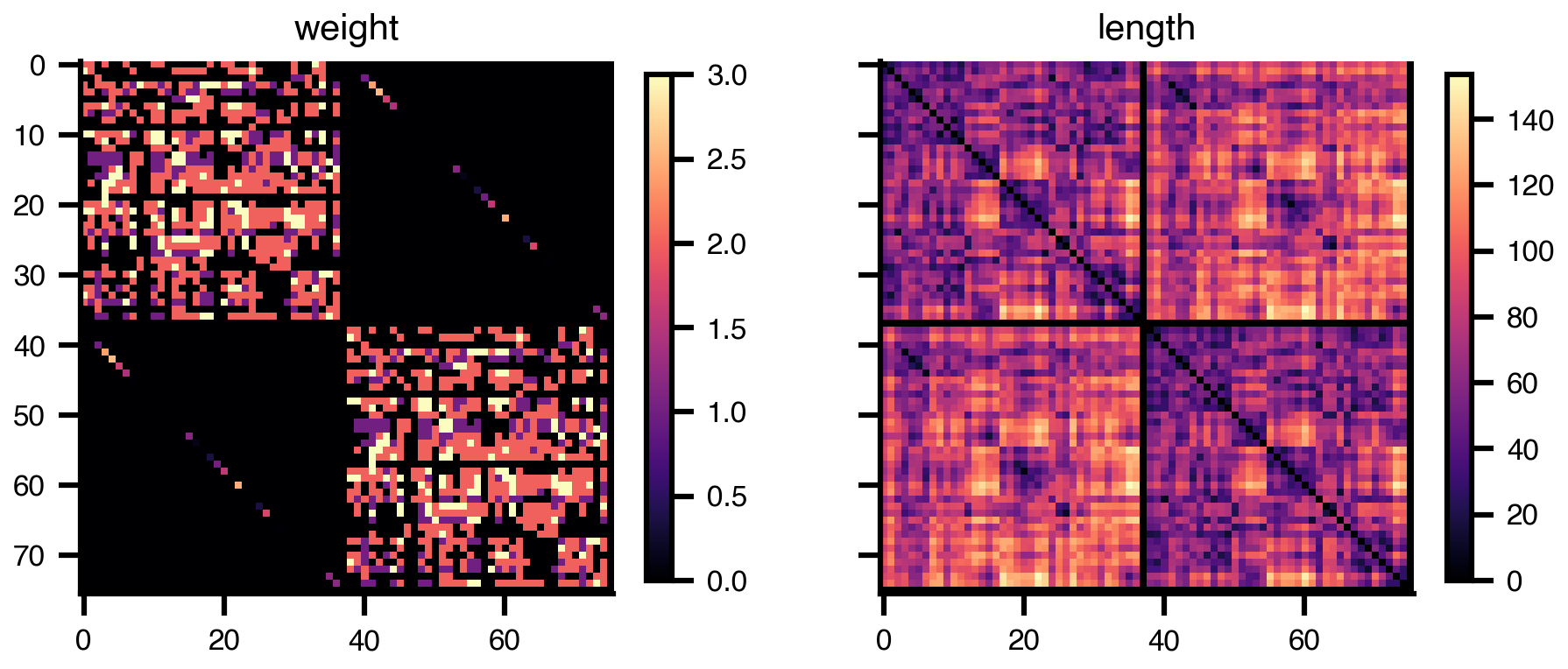
net.plot_overview(plot_brain=False)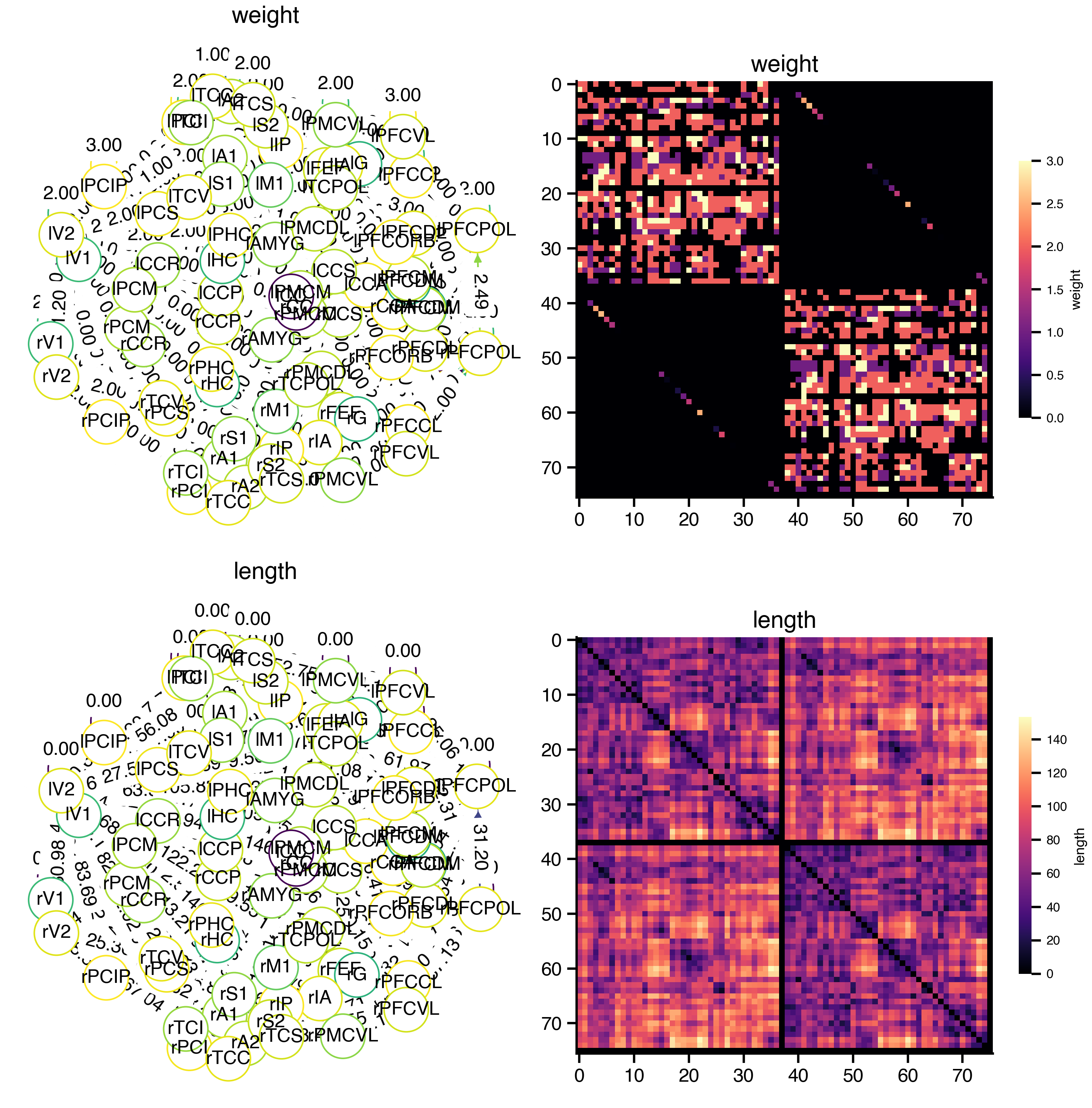
3. Save as YAML + HDF5
TVBO stores networks as a human-readable YAML sidecar (metadata and schema) plus an HDF5 companion file (dense matrix data with gzip compression).
outdir = Path(tempfile.mkdtemp())
yaml_path = outdir / "tvb_default_76.yaml"
net.save(yaml_path)
print("Created files:")
for f in sorted(outdir.iterdir()):
sz = f.stat().st_size
print(f" {f.name:30s} ({sz:,} bytes)")Created files:
tvb_default_76.h5 (64,256 bytes)
tvb_default_76.yaml (15,549 bytes)The YAML sidecar is fully self-describing:
# Show first 40 lines of the sidecar
lines = yaml_path.read_text().splitlines()
for line in lines[:40]:
print(line)
print(f"... ({len(lines)} lines total)")tvbo_class: tvbo:Network
schema_version: tvb-datamodel/0.7.0
label: 'Connectivity gid: 9df860e7-5404-4cf5-98d7-500e11dfbbed'
number_of_nodes: 76
descriptor: SC
distance_unit: mm
time_unit: ms
data_file: tvb_default_76.h5
parameters:
conduction_speed:
label: v
value: 3.0
unit: mm_per_ms
nodes:
- id: 0
label: rA1
parameters:
cortical:
value: 1.0
area:
value: 396.44065
hemisphere:
value: 1.0
position:
x: -9.885591
y: -47.084818
z: -3.13936
- id: 1
label: rA2
parameters:
cortical:
value: 1.0
area:
value: 937.50289
hemisphere:
value: 1.0
position:
x: -2.605247
y: -55.324507
z: -7.065423
... (1015 lines total)4. Reload and Verify
reloaded = Network.from_file(yaml_path)
print(f"Reloaded: {reloaded.number_of_nodes} nodes")
print(f"Label: {reloaded.label}")Reloaded: 76 nodes
Label: Connectivity gid: 9df860e7-5404-4cf5-98d7-500e11dfbbedVerify lossless round-trip — arrays match exactly:
np.testing.assert_allclose(reloaded.weights_matrix, net.weights_matrix)
np.testing.assert_allclose(reloaded.lengths_matrix, net.lengths_matrix)
print("✓ Weights match")
print("✓ Lengths match")✓ Weights match
✓ Lengths matchNode metadata also survives:
for i in range(min(5, net.number_of_nodes)):
orig = net.nodes[i]
load = reloaded.nodes[i]
assert orig.label == load.label, f"Label mismatch at node {i}"
assert abs(orig.position.x - load.position.x) < 1e-6
print(f"✓ First {min(5, net.number_of_nodes)} node labels and positions match")✓ First 5 node labels and positions match5. Export Back to TVB
Convert the reloaded network back to a TVB Connectivity and compare with the original:
conn2 = reloaded.execute(format="tvb")
print(f"TVB round-trip: {conn2.weights.shape[0]} regions")
np.testing.assert_allclose(conn2.weights, conn.weights)
np.testing.assert_allclose(conn2.tract_lengths, conn.tract_lengths)
np.testing.assert_allclose(conn2.centres, conn.centres, atol=1e-5)
np.testing.assert_allclose(conn2.speed, conn.speed)
print("✓ weights identical")
print("✓ tract_lengths identical")
print("✓ centres identical (atol=1e-5)")
print(f"✓ speed identical ({conn2.speed[0]} mm/ms)")TVB round-trip: 76 regions
✓ weights identical
✓ tract_lengths identical
✓ centres identical (atol=1e-5)
✓ speed identical (3.0 mm/ms)Centre coordinates
import matplotlib.pyplot as plt
fig, axes = plt.subplots(1, 3, figsize=(12, 3.5))
for ax, (dim, lbl) in zip(axes, enumerate(["x (mm)", "y (mm)", "z (mm)"])):
orig = conn.centres[:, dim]
rt = conn2.centres[:, dim]
ax.scatter(orig, rt, s=8, alpha=0.7)
lims = [min(orig.min(), rt.min()) - 2, max(orig.max(), rt.max()) + 2]
ax.plot(lims, lims, 'k--', lw=0.8, alpha=0.5)
ax.set_xlim(lims); ax.set_ylim(lims)
ax.set_xlabel(f"Original {lbl}"); ax.set_ylabel(f"Round-trip {lbl}")
ax.set_aspect('equal')
fig.suptitle("Centre coordinates: original vs round-trip", fontsize=11)
plt.tight_layout()
plt.show()
print(f"Max centre deviation: {np.abs(conn2.centres - conn.centres).max():.2e} mm")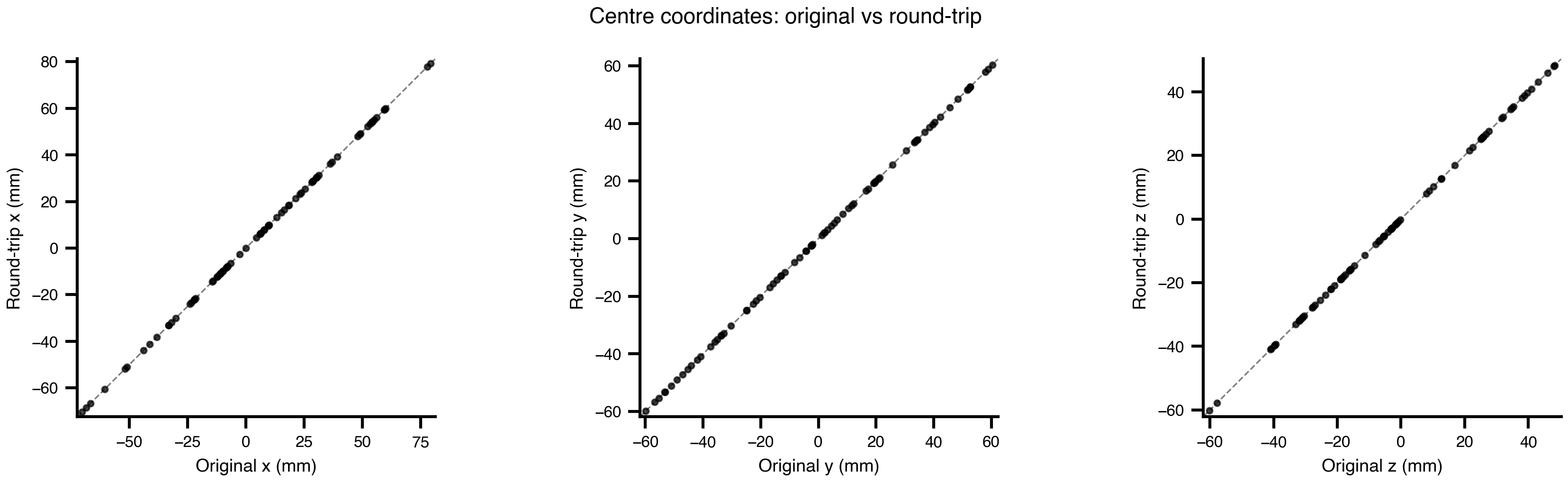
Max centre deviation: 0.00e+00 mmdiff = conn2.weights - conn.weights
fig, ax = plt.subplots(figsize=(6, 5))
im = ax.imshow(diff, cmap="RdBu_r", vmin=-1e-10, vmax=1e-10)
ax.set_title("Weight difference (original − round-trip)")
ax.set_xlabel("Region")
ax.set_ylabel("Region")
fig.colorbar(im, ax=ax, label="Δweight")
plt.tight_layout()
plt.show()
print(f"Max absolute difference: {np.abs(diff).max():.2e}")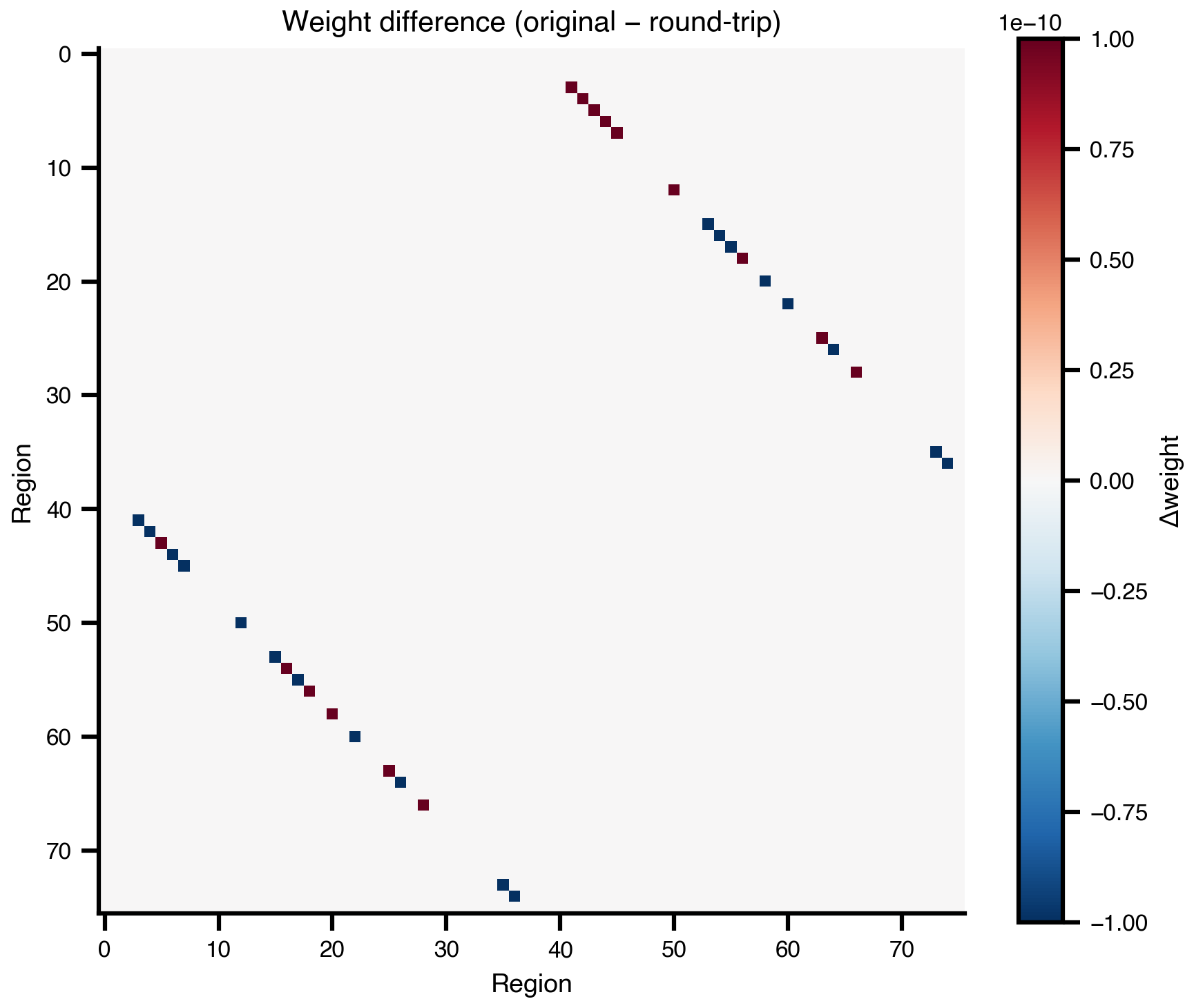
Max absolute difference: 1.18e-076. Surface Network (Multi-Level)
TVB surface simulations use three objects: Connectivity (regions), CorticalSurface (mesh), and RegionMapping (vertex → region). TVBO imports all three into a linked pair of networks.
from tvb.datatypes.surfaces import CorticalSurface
from tvb.datatypes.region_mapping import RegionMapping
surf = CorticalSurface.from_file()
surf.configure()
rmap = RegionMapping.from_file()
rmap.connectivity = conn
rmap.surface = surf
rmap.configure()
print(f"Surface: {surf.vertices.shape[0]} vertices, "
f"{surf.triangles.shape[0]} triangles")
print(f"Region mapping: {rmap.array_data.shape[0]} entries → "
f"{len(np.unique(rmap.array_data))} regions")Surface: 16384 vertices, 32760 triangles
Region mapping: 16384 entries → 76 regionsregion_net, surface_net = Network.from_tvb_surface(conn, surf, rmap)
print(f"Region network: {region_net.number_of_nodes} nodes")
print(f"Surface network: {surface_net.number_of_nodes} nodes")
print(f"Mesh: {surface_net.mesh.number_of_vertices} vertices, "
f"{surface_net.mesh.number_of_elements} triangles")
print(f"Parent link: {surface_net.parent_network is not None}")Region network: 76 nodes
Surface network: 16384 nodes
Mesh: 16384 vertices, 32760 triangles
Parent link: TrueVisualize the Cortical Mesh
vertices = surface_net._mesh_vertices
triangles = surface_net._mesh_elements
region_ids = surface_net.node_mapping_data.astype(float)
fig, axes = plt.subplots(1, 2, figsize=(10, 5))
plot_surf(
vertices=vertices, faces=triangles, overlay=region_ids,
view='lateral', cmap='tab20', parcellated=True, ax=axes[0],
)
axes[0].set_title("Lateral")
plot_surf(
vertices=vertices, faces=triangles, overlay=region_ids,
view='dorsal', cmap='tab20', parcellated=True, ax=axes[1],
)
axes[1].set_title("Dorsal")
plt.tight_layout()
plt.show()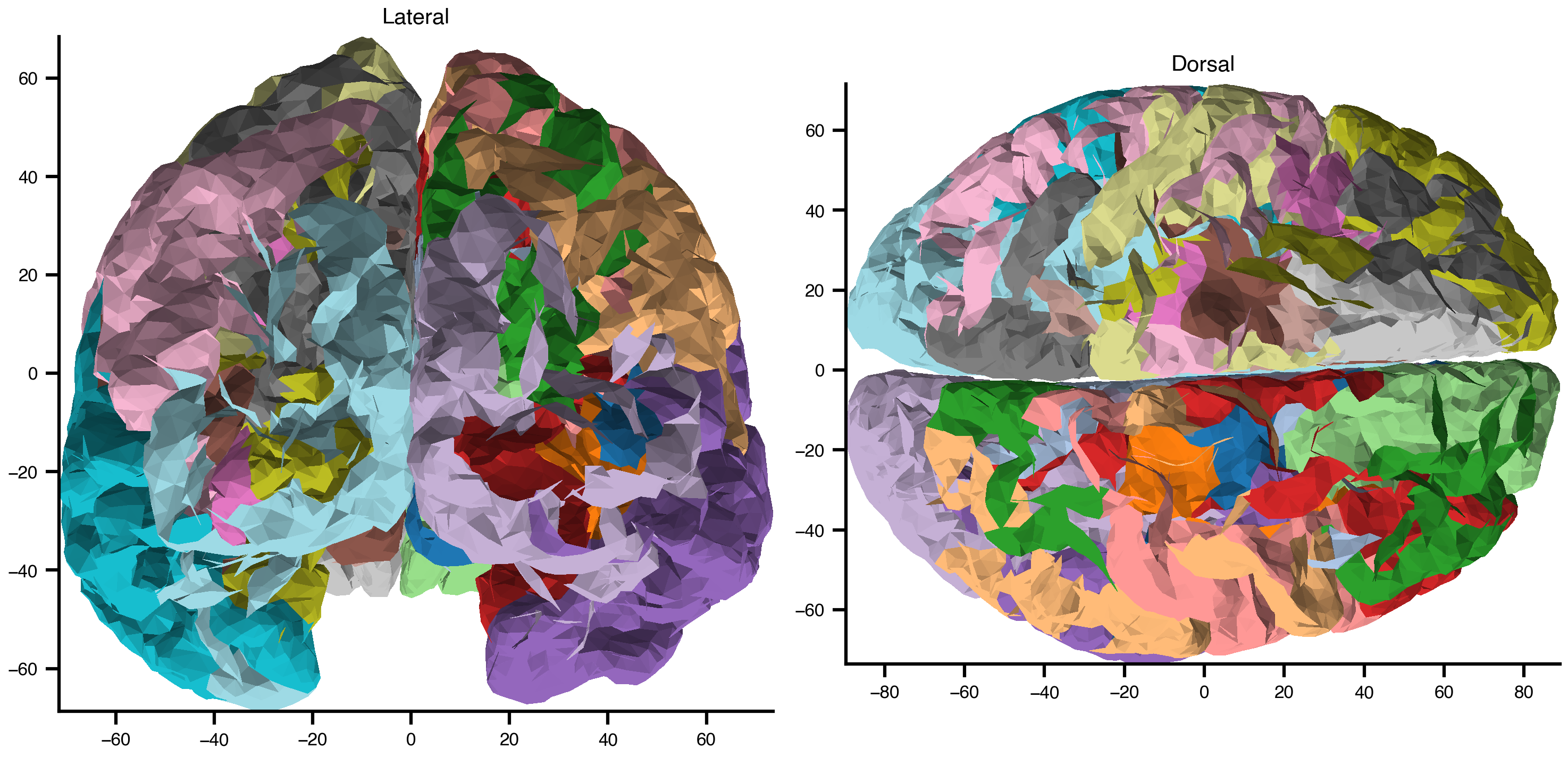
Save and Reload Multi-Level Networks
surf_dir = Path(tempfile.mkdtemp())
region_net.save(surf_dir / "regions.yaml")
surface_net.save(surf_dir / "surface.yaml")
print("Surface network files:")
for f in sorted(surf_dir.iterdir()):
sz = f.stat().st_size
print(f" {f.name:30s} ({sz:,} bytes)")Surface network files:
regions.h5 (64,256 bytes)
regions.yaml (15,542 bytes)
surface.h5 (823,061 bytes)
surface.yaml (2,002,533 bytes)reloaded_surf = Network.from_file(surf_dir / "surface.yaml")
print(f"Reloaded: {reloaded_surf.number_of_nodes} nodes")
print(f"Mesh vertices: {reloaded_surf._mesh_vertices.shape}")
print(f"Mesh triangles: {reloaded_surf._mesh_elements.shape}")
np.testing.assert_allclose(reloaded_surf._mesh_vertices, vertices)
np.testing.assert_allclose(reloaded_surf._mesh_elements, triangles)
print("✓ Mesh data survives HDF5 round-trip")Reloaded: 16384 nodes
Mesh vertices: (16384, 3)
Mesh triangles: (32760, 3)
✓ Mesh data survives HDF5 round-tripOptional: MNE Projection Layer in the Network Section
The MNE exporter creates a sensor-to-region projection network in tvbo format (sensors_eeg_standard1005_fsaverage_aparc_projection.yaml). This can be handled in the same multi-layer network workflow as other network resources.
from tvbo import database_path
mne_yaml = (
database_path
/ "networks"
/ "sensors_eeg_standard1005_fsaverage_aparc_projection.yaml"
)
mne_meta = mne_yaml.with_name(f"{mne_yaml.stem}_plotmeta.npz")
if mne_yaml.exists() and mne_meta.exists():
mne_net = Network.from_file(mne_yaml)
mne_gain = np.asarray(mne_net.matrix("gain", format="dense"), dtype=float)
mne_plotmeta = np.load(mne_meta)
mne_region_labels = [str(x) for x in mne_plotmeta["region_labels"]]
mne_region_centers = np.asarray(mne_plotmeta["region_centers_mm"], dtype=float)
mne_sensor_labels = [str(node.label) for node in mne_net.nodes]
mne_sensor_pos = np.asarray(
[[n.position.x, n.position.y, n.position.z] for n in mne_net.nodes],
dtype=float,
)
n_regions = len(mne_region_labels)
n_sensors = len(mne_sensor_labels)
n_total = n_regions + n_sensors
centers = {}
for i, lbl in enumerate(mne_region_labels):
centers[f"R:{lbl}"] = tuple(mne_region_centers[i])
for i, lbl in enumerate(mne_sensor_labels):
centers[f"S:{lbl}"] = tuple(mne_sensor_pos[i])
combined = np.zeros((n_total, n_total), dtype=float)
for i in range(n_sensors):
for j in range(n_regions):
combined[n_regions + i, j] = abs(mne_gain[i, j])
labels = [f"R:{x}" for x in mne_region_labels] + [f"S:{x}" for x in mne_sensor_labels]
mne_graph = create_network(
centers,
{"mne_forward_gain": combined},
labels=labels,
threshold_percentile=80,
directed=True,
edge_data_key="gain",
)
print(
f"MNE projection layer: nodes={mne_graph.number_of_nodes()}, "
f"edges={mne_graph.number_of_edges()}, gain_shape={mne_gain.shape}"
)
else:
print(
"MNE projection network not found. Generate it with:\n"
"python scripts/export_mne_standard1005_projection_network.py"
)MNE projection network not found. Generate it with:
python scripts/export_mne_standard1005_projection_network.pySummary
| Step | Method | What It Does |
|---|---|---|
| Import | Network.from_tvb(conn) |
TVB Connectivity → tvbo Network |
| Visualize | net.plot_matrix(), net.plot_overview() |
Weight/length heatmaps, graph layout |
| Save | net.save("net.yaml") |
YAML sidecar + HDF5 companion |
| Load | Network.from_file("net.yaml") |
Lazy load from file pair |
| Export | net.execute(format="tvb") |
tvbo Network → TVB Connectivity |
| Surface | Network.from_tvb_surface(conn, surf, rmap) |
Multi-level: regions + mesh |
The full pipeline is lossless — all matrices, node metadata, mesh geometry, and region mappings survive the round-trip with zero numerical error.
