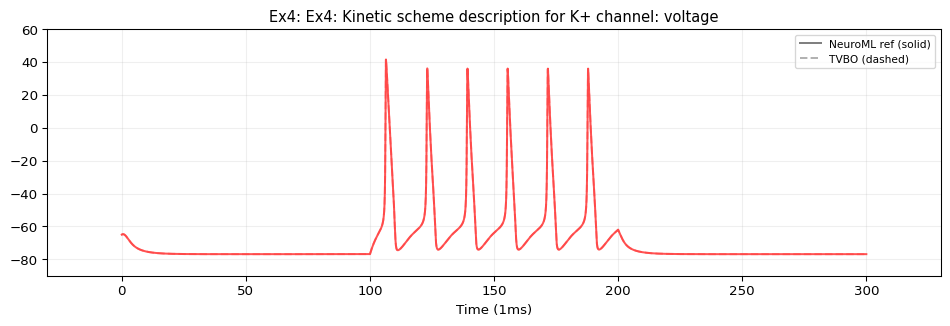
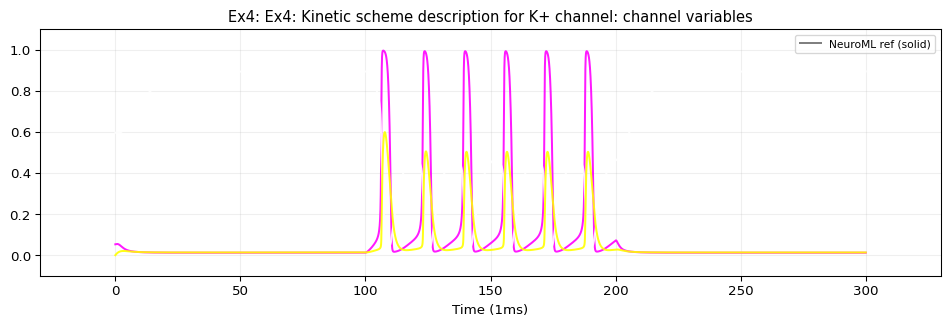
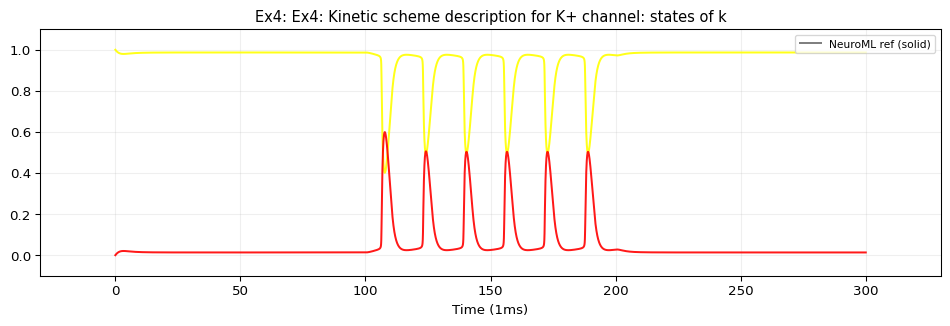
Model: Kinetic Scheme Ion Channels
This example demonstrates ionChannelKS — ion channels defined by Markov state transitions between open/closed states. The k_vh channel uses a vHalfTransition with rates:
\[r_{f0} = \frac{\exp(z\gamma(v - v_{Half})/k_{te})}{\tau}, \quad
r_{r0} = \frac{\exp(-z(1-\gamma)(v - v_{Half})/k_{te})}{\tau}\]
\[r_f = \frac{1}{1/r_{f0} + \tau_{Min}}, \quad
r_r = \frac{1}{1/r_{r0} + \tau_{Min}}\]
where \(k_{te} = 25.3\,\text{mV}\) . The cell has a standard HH Na channel (\(m^3 h\) ) and the KS K channel (\(n\) , instances=1):
\[C\frac{dv}{dt} = -g_{Na}m^3 h(v - E_{Na}) - g_K n(v - E_K) + I_{\text{ext}}\]
By annotating the dynamics with iri: neuroml:pointCellCondBased and structuring the ion channels as components (including ionChannelKS with gateKS and vHalfTransition), TVBO emits standard NeuroML2 types that preserve the jLEMS child-before-parent RK4 evaluation order — giving exact numerical identity.
1. Define in TVBO
from tvbo import SimulationExperiment# Ex4 parameters: pointCellCondBased, C=1pF # Na: 6000 channels × 20pS, erev=50mV (standard HH rates) # K (k_vh): 1800 channels × 8pS, erev=-77mV (KS vHalfTransition) = SimulationExperiment.from_string(""" label: "NeuroML Ex4: KS Channels" dynamics: name: HH_KineticScheme iri: neuroml:pointCellCondBased description: > Cell with HH Na channel and kinetic-scheme K channel, matching NeuroML2 Ex4. parameters: C: { value: 1, unit: pF } v0: { value: -65, unit: mV } thresh: { value: -20, unit: mV } pulse_delay: { value: 100, unit: ms } pulse_duration: { value: 100, unit: ms } I_amp: { value: 0.005, unit: nA } components: na: name: na iri: neuroml:ionChannelHH parameters: conductance: { value: 20, unit: pS } number: { value: 6000 } erev: { value: 50, unit: mV } components: m: name: m iri: neuroml:gateHHrates parameters: instances: { value: 3 } components: forwardRate: name: forwardRate iri: neuroml:HHExpLinearRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -40, unit: mV } scale: { value: 10, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHExpRate parameters: rate: { value: 4, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -18, unit: mV } h: name: h iri: neuroml:gateHHrates parameters: instances: { value: 1 } components: forwardRate: name: forwardRate iri: neuroml:HHExpRate parameters: rate: { value: 0.07, unit: per_ms } midpoint: { value: -65, unit: mV } scale: { value: -20, unit: mV } reverseRate: name: reverseRate iri: neuroml:HHSigmoidRate parameters: rate: { value: 1, unit: per_ms } midpoint: { value: -35, unit: mV } scale: { value: 10, unit: mV } k_vh: name: k_vh iri: neuroml:ionChannelKS parameters: conductance: { value: 8, unit: pS } number: { value: 1800 } erev: { value: -77, unit: mV } components: n: name: n iri: neuroml:gateKS parameters: instances: { value: 1 } components: c1: name: c1 iri: neuroml:closedState o1: name: o1 iri: neuroml:openState vh1: name: vh1 iri: neuroml:vHalfTransition parameters: from: { description: c1 } to: { description: o1 } vHalf: { value: 0, unit: mV } z: { value: 1.5 } gamma: { value: 0.75 } tau: { value: 3.2, unit: ms } tauMin: { value: 0.3, unit: ms } integration: step_size: 0.01 duration: 300.0 time_scale: ms """ )print (f"Model: { exp. dynamics. name if exp. dynamics else 'network' } " )print (f"IRI: { exp. dynamics. iri} " )print (f"Components: { list (exp.dynamics.components.keys())} " )
Model: HH_KineticScheme
IRI: neuroml:pointCellCondBased
Components: ['k_vh', 'na']
This example uses iri: neuroml:ionChannelKS with gateKS containing closedState, openState, and vHalfTransition sub-components. TVBO’s adapter emits these as standard NeuroML2 XML elements — no need to flatten the Markov scheme into ODEs. This gives exact numerical identity with the reference simulation at the original step size (0.01 ms).
2. Render LEMS XML
= exp.render("lems" )for line in xml.split(' \n ' ):= line.strip()if any (tag in line_stripped for tag in ['<ionChannel' , '<gateHH' , '<gateKS' ,'<pointCell' , '<channelPopulation' , '<closedState' , '<openState' ,'<vHalfTransition' , '<pulseGenerator' , 'Rate type=' , '<Include' ]):print (line_stripped)
<Include file="Cells.xml"/>
<Include file="Networks.xml"/>
<Include file="Inputs.xml"/>
<Include file="Simulation.xml"/>
<ionChannelKS id="k_vh" conductance="8 pS">
<gateKS id="n" instances="1">
<closedState id="c1"/>
<openState id="o1"/>
<vHalfTransition id="vh1" from="c1" gamma="0.75" tau="3.2 ms" tauMin="0.3 ms" to="o1" vHalf="0 mV" z="1.5"/>
<ionChannelHH id="na" conductance="20 pS">
<gateHHrates id="h" instances="1">
<forwardRate type="HHExpRate" midpoint="-65 mV" rate="0.07 per_ms" scale="-20 mV"/>
<reverseRate type="HHSigmoidRate" midpoint="-35 mV" rate="1 per_ms" scale="10 mV"/>
<gateHHrates id="m" instances="3">
<forwardRate type="HHExpLinearRate" midpoint="-40 mV" rate="1 per_ms" scale="10 mV"/>
<reverseRate type="HHExpRate" midpoint="-65 mV" rate="4 per_ms" scale="-18 mV"/>
<pointCellCondBased id="HH_KineticScheme" C="1 pF" v0="-65 mV" thresh="-20 mV">
<channelPopulation id="k_vh_pop" ionChannel="k_vh" number="1800" erev="-77 mV"/>
<channelPopulation id="na_pop" ionChannel="na" number="6000" erev="50 mV"/>
<pulseGenerator id="pulseGen1" delay="100 ms" duration="100 ms" amplitude="0.005 nA"/>
3. Run Reference
from tvbo.adapters.neuroml import run_lems_example= run_lems_example("LEMS_NML2_Ex4_KS.xml" )for name, arr in ref_outputs.items():print (f" { name} : shape= { arr. shape} " )
auto.dat: shape=(30001, 7)
4. Run TVBO Version
= exp.run("neuroml" )= result.integration.dataprint (f"TVBO: { da. dims} , shape= { da. shape} " )
TVBO: ('time', 'variable'), shape=(30001, 1)
5. Compare & Plot
from tvbo.adapters.neuroml import plot_lems_comparison"LEMS_NML2_Ex4_KS.xml" , ref_outputs, result.integration.data, title_prefix= "Ex4" )